Pharmacogenetic landscape of Metabolic Syndrome components drug response in Tunisia and comparison with worldwide populations
- PMID: 29652911
- PMCID: PMC5898725
- DOI: 10.1371/journal.pone.0194842
Pharmacogenetic landscape of Metabolic Syndrome components drug response in Tunisia and comparison with worldwide populations
Abstract
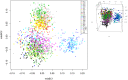
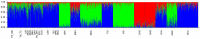
Genetic variation is an important determinant affecting either drug response or susceptibility to adverse drug reactions. Several studies have highlighted the importance of ethnicity in influencing drug response variability that should be considered during drug development. Our objective is to characterize the genetic variability of some pharmacogenes involved in the response to drugs used for the treatment of Metabolic Syndrome (MetS) in Tunisia and to compare our results to the worldwide populations. A set of 135 Tunisians was genotyped using the Affymetrix Chip 6.0 genotyping array. Variants located in 24 Very Important Pharmacogenes (VIP) involved in MetS drug response were extracted from the genotyping data. Analysis of variant distribution in Tunisian population compared to 20 worldwide populations publicly available was performed using R software packages. Common variants between Tunisians and the 20 investigated populations were extracted from genotyping data. Multidimensional screening showed that Tunisian population is clustered with North African and European populations. The greatest divergence was observed with the African and Asian population. In addition, we performed Inter-ethnic comparison based on the genotype frequencies of five VIP biomarkers. The genotype frequencies of the biomarkers rs3846662, rs1045642, rs7294 and rs12255372 located respectively in HMGCR, ABCB1, VKORC1 and TCF7L2 are similar between Tunisian, Tuscan (TSI) and European (CEU). The genotype frequency of the variant rs776746 located in CYP3A5 gene is similar between Tunisian and African populations and different from CEU and TSI. The present study shows that the genetic make up of the Tunisian population is relatively complex in regard to pharmacogenes and reflects previous historical events. It is important to consider this ethnic difference in drug prescription in order to optimize drug response to avoid serious adverse drug reactions. Taking into account similarities with other neighboring populations, our study has an impact not only on the Tunisian population but also on North African population which are underrepresented in pharmacogenomic studies.
Conflict of interest statement
Figures
References
-
- Wang L, Aikemu A, Yibulayin A, Du S, Geng T, Wang B, et al. Genetic polymorphisms of pharmacogenomic VIP variants in the Uygur population from northwestern China. BMC genetics. 2015;16:66 Epub 2015/06/21. doi: 10.1186/s12863-015-0232-x . - DOI - PMC - PubMed
-
- Pirmohamed M. Personalized pharmacogenomics: predicting efficacy and adverse drug reactions. Annual review of genomics and human genetics. 2014;15:349–70. Epub 2014/06/06. doi: 10.1146/annurev-genom-090413-025419 . - DOI - PubMed
-
- Weinshilboum R. Inheritance and drug response. The New England journal of medicine. 2003;348(6):529–37. Epub 2003/02/07. doi: 10.1056/NEJMra020021 . - DOI - PubMed
-
- Johnson JA. Pharmacogenetics: potential for individualized drug therapy through genetics. Trends in genetics: TIG. 2003;19(11):660–6. Epub 2003/10/31. doi: 10.1016/j.tig.2003.09.008 . - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

