A TAD boundary is preserved upon deletion of the CTCF-rich Firre locus
- PMID: 29654311
- PMCID: PMC5899154
- DOI: 10.1038/s41467-018-03614-0
A TAD boundary is preserved upon deletion of the CTCF-rich Firre locus
Abstract
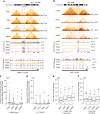
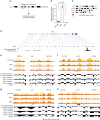
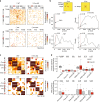
The binding of the transcriptional regulator CTCF to the genome has been implicated in the formation of topologically associated domains (TADs). However, the general mechanisms of folding the genome into TADs are not fully understood. Here we test the effects of deleting a CTCF-rich locus on TAD boundary formation. Using genome-wide chromosome conformation capture (Hi-C), we focus on one TAD boundary on chromosome X harboring ~ 15 CTCF binding sites and located at the long non-coding RNA (lncRNA) locus Firre. Specifically, this TAD boundary is invariant across evolution, tissues, and temporal dynamics of X-chromosome inactivation. We demonstrate that neither the deletion of this locus nor the ectopic insertion of Firre cDNA or its ectopic expression are sufficient to alter TADs in a sex-specific or allele-specific manner. In contrast, Firre's deletion disrupts the chromatin super-loop formation of the inactive X-chromosome. Collectively, our findings suggest that apart from CTCF binding, additional mechanisms may play roles in establishing TAD boundary formation.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Structural organization of the inactive X chromosome in the mouse.Nature. 2016 Jul 28;535(7613):575-9. doi: 10.1038/nature18589. Epub 2016 Jul 18. Nature. 2016. PMID: 27437574 Free PMC article.
-
The lncRNA Firre anchors the inactive X chromosome to the nucleolus by binding CTCF and maintains H3K27me3 methylation.Genome Biol. 2015 Mar 12;16(1):52. doi: 10.1186/s13059-015-0618-0. Genome Biol. 2015. PMID: 25887447 Free PMC article.
-
HOTTIP-dependent R-loop formation regulates CTCF boundary activity and TAD integrity in leukemia.Mol Cell. 2022 Feb 17;82(4):833-851.e11. doi: 10.1016/j.molcel.2022.01.014. Mol Cell. 2022. PMID: 35180428 Free PMC article.
-
Pushing the TAD boundary: Decoding insulator codes of clustered CTCF sites in 3D genomes.Bioessays. 2024 Oct;46(10):e2400121. doi: 10.1002/bies.202400121. Epub 2024 Aug 21. Bioessays. 2024. PMID: 39169755 Review.
-
Multifaceted role of CTCF in X-chromosome inactivation.Chromosoma. 2024 Oct;133(4):217-231. doi: 10.1007/s00412-024-00826-w. Epub 2024 Oct 21. Chromosoma. 2024. PMID: 39433641 Review.
Cited by
-
Differential contribution of steady-state RNA and active transcription in chromatin organization.EMBO Rep. 2019 Oct 4;20(10):e48068. doi: 10.15252/embr.201948068. Epub 2019 Aug 26. EMBO Rep. 2019. PMID: 31448565 Free PMC article.
-
Outward-oriented sites within clustered CTCF boundaries are key for intra-TAD chromatin interactions and gene regulation.Nat Commun. 2023 Dec 7;14(1):8101. doi: 10.1038/s41467-023-43849-0. Nat Commun. 2023. PMID: 38062010 Free PMC article.
-
Engineering three-dimensional genome folding.Nat Genet. 2021 May;53(5):602-611. doi: 10.1038/s41588-021-00860-9. Epub 2021 May 6. Nat Genet. 2021. PMID: 33958782 Review.
-
A global high-density chromatin interaction network reveals functional long-range and trans-chromosomal relationships.Genome Biol. 2022 Nov 9;23(1):238. doi: 10.1186/s13059-022-02790-z. Genome Biol. 2022. PMID: 36352464 Free PMC article.
-
Mini-review: Gene regulatory network benefits from three-dimensional chromatin conformation and structural biology.Comput Struct Biotechnol J. 2023 Feb 16;21:1728-1737. doi: 10.1016/j.csbj.2023.02.028. eCollection 2023. Comput Struct Biotechnol J. 2023. PMID: 36890880 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

