Genome-wide identification of long non-coding RNAs suggests a potential association with effector gene transcription in Phytophthora sojae
- PMID: 29665235
- PMCID: PMC6638102
- DOI: 10.1111/mpp.12692
Genome-wide identification of long non-coding RNAs suggests a potential association with effector gene transcription in Phytophthora sojae
Abstract
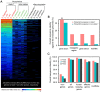
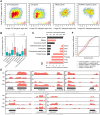
Numerous long non-coding RNAs (lncRNAs) identified and characterized in mammals, plants and fungi have been found to play critical regulatory roles in biological processes. However, little is known about the role of lncRNAs in oomycete plant pathogens, which cause devastating damage to the economy and ecosystems. We used strand-specific RNA sequencing (RNA-seq) to generate a computational pipeline to identify lncRNAs in Phytophthora sojae, a model oomycete plant pathogen. In total, 940 lncRNAs with 1010 isoforms were identified from RNA-seq data obtained from four representative stages of P. sojae. The lncRNAs had shorter transcript lengths, longer exon lengths, fewer numbers of exons, lower GC content and higher minimum free energy values compared with protein-coding genes. lncRNAs in P. sojae exhibited low sequence conservation amongst oomycetes and P. sojae isolates. Transcriptional data indicated that P. sojae lncRNAs tended to be transcribed in a stage-specific manner; representative lncRNAs were validated by semi-quantitative reverse transcription-polymerase chain reaction. Phytophthora sojae lncRNAs were concentrated in gene-sparse regions, and lncRNAs were associated with secreted protein and effector coding genes. The neighbouring genes of lncRNAs encoded various effector family members, and RNA-seq data revealed a correlation between the transcription level of lncRNAs and their neighbouring genes. Our results provide the first comprehensive identification of lncRNAs in oomycetes and suggest a potential association between lncRNAs and effector genes.
Keywords: Phytophthora; RNA-seq; effector; long non-coding RNAs; transcriptional regulation.
© 2018 BSPP and John Wiley & Sons Ltd.
Figures


References
-
- Asman, A.K. , Fogelqvist, J. , Vetukuri, R.R. and Dixelius, C. (2016) Phytophthora infestans argonaute 1 binds microRNA and small RNAs from effector genes and transposable elements. New Phytol. 211, 993–1007. - PubMed
-
- Avrova, A.O. , Whisson, S.C. , Pritchard, L. , Venter, E. , De Luca, S. , Hein, I. and Birch, P.R.J. (2007) A novel non‐protein‐coding infection‐specific gene family is clustered throughout the genome of Phytophthora infestans . Microbiology, 153, 747–759. - PubMed
-
- Baldauf, S.L. (2003) The deep roots of eukaryotes. Science, 300, 1703–1706. - PubMed
-
- Baxter, L. , Tripathy, S. , Ishaque, N. , Boot, N. , Cabral, A. , Kemen, E. , Thines, M. , Ah‐Fong, A. , Anderson, R. , Badejoko, W. , Bittner‐Eddy, P. , Boore, J.L. , Chibucos, M.C. , Coates, M. , Dehal, P. , Delehaunty, K. , Dong, S. , Downton, P. , Dumas, B. , Fabro, G. , Fronick, C. , Fuerstenberg, S.I. , Fulton, L. , Gaulin, E. , Govers, F. , Hughes, L. , Humphray, S. , Jiang, R.H.Y. , Judelson, H. , Kamoun, S. , Kyung, K. , Meijer, H. , Minx, P. , Morris, P. , Nelson, J. , Phuntumart, V. , Qutob, D. , Rehmany, A. , Rougon‐Cardoso, A. , Ryden, P. , Torto‐Alalibo, T. , Studholme, D. , Wang, Y. , Win, J. , Wood, J. , Clifton, S.W. , Rogers, J. , Van den Ackerveken, G. , Jones, J.D.G. , McDowell, J.M. , Beynon, J. and Tyler, B.M. (2010) Signatures of adaptation to obligate biotrophy in the Hyaloperonospora arabidopsidis genome. Science, 330, 1549–1551. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

