Serotype Diversity and Antimicrobial Resistance among Salmonella enterica Isolates from Patients at an Equine Referral Hospital
- PMID: 29678910
- PMCID: PMC6007101
- DOI: 10.1128/AEM.02829-17
Serotype Diversity and Antimicrobial Resistance among Salmonella enterica Isolates from Patients at an Equine Referral Hospital
Abstract
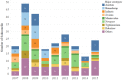
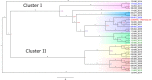
Although Salmonella enterica can produce life-threatening colitis in horses, certain serotypes are more commonly associated with clinical disease. Our aim was to evaluate the proportional morbidity attributed to different serotypes, as well as the phenotypic and genotypic antimicrobial resistance (AMR) of Salmonella isolates from patients at an equine referral hospital in the southern United States. A total of 255 Salmonella isolates was obtained from clinical samples of patients admitted to the hospital between 2007 and 2015. Phenotypic resistance to 14 antibiotics surveilled by the U.S. National Antimicrobial Resistance Monitoring System was determined using a commercially available panel. Whole-genome sequencing was used to identify serotypes and genotypic AMR. The most common serotypes were Salmonella enterica serotype Newport (18%), Salmonella enterica serotype Anatum (15.2%), and Salmonella enterica serotype Braenderup (11.8%). Most (n = 219) of the isolates were pansusceptible, while 25 were multidrug resistant (≥3 antimicrobial classes). Genes encoding beta-lactam resistance, such as blaCMY-2, blaSHV-12, blaCTX-M-27, and blaTEM-1B, were detected. The qnrB2 and aac(6')-Ib-cr genes were present in isolates with reduced susceptibility to ciprofloxacin. Genes encoding resistance to gentamicin (aph(3')-Ia, aac(6')-IIc), streptomycin (strA and strB), sulfonamides (sul1), trimethoprim (dfrA), phenicols (catA), tetracyclines [tet(A) and tet(E)], and macrolides [ere(A)] were also identified. The main predicted incompatibility plasmid type was I1 (10%). Core genome-based analyses revealed phylogenetic associations between isolates of common serotypes. The presence of AMR Salmonella in equine patients increases the risk of unsuccessful treatment and causes concern for potential zoonotic transmission to attending veterinary personnel, animal caretakers, and horse owners. Understanding the epidemiology of Salmonella in horses admitted to referral hospitals is important for the prevention, control, and treatment of salmonellosis.IMPORTANCE In horses, salmonellosis is a leading cause of life-threatening colitis. At veterinary teaching hospitals, nosocomial outbreaks can increase the risk of zoonotic transmission, lead to restrictions on admissions, impact hospital reputation, and interrupt educational activities. The antimicrobials most often used in horses are included in the 5th revision of the World Health Organization's list of critically important antimicrobials for human medicine. Recent studies have demonstrated a trend of increasing bacterial resistance to drugs commonly used to treat Salmonella infections. In this study, we identify temporal trends in the distribution of Salmonella serotypes and their mechanisms of antimicrobial resistance; furthermore, we are able to determine the likely origin of several temporal clusters of infection by using whole-genome sequencing. These data can be used to focus strategies to better contain the dissemination and enhance the mitigation of Salmonella infections and to provide evidence-based policies and guidelines to steward antimicrobial use in veterinary medicine.
Keywords: Salmonella; antibiotic resistance; antimicrobial agents; horses; nosocomial; whole-genome sequencing.
Copyright © 2018 American Society for Microbiology.
Figures


References
-
- Chapman AM. 2006. Characterizing Salmonella fecal shedding among racehorses in Louisiana. MSc thesis Louisiana State University, Baton Rouge, LA.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

