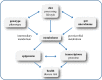
Impact of dietary gut microbial metabolites on the epigenome
- PMID: 29685968
- PMCID: PMC5915727
- DOI: 10.1098/rstb.2017.0359
Impact of dietary gut microbial metabolites on the epigenome
Abstract
Within the past decade, epigenetic mechanisms and their modulation by natural products have gained increasing interest. Dietary bioactive compounds from various sources, including green tea, soya, fruit and berries, cruciferous vegetables, whole grain foods, fish and others, have been shown to target enzymes involved in epigenetic gene regulation, including DNA methyltransferases, histone acetyltransferases, deacetylases and demethylases in vitro and in cell culture. Also, many dietary agents were shown to alter miRNA expression. In vivo studies in animal models and humans are still limited. Recent research has indicated that the gut microbiota and gut microbial metabolites might be important mediators of diet-epigenome interactions. Inter-individual differences in the gut microbiome might affect release, metabolism and bioavailability of dietary agents and explain variability in response to intervention in human studies. Only a few microbial metabolites, including folate, phenolic acids, S-(-)equol, urolithins, isothiocyanates, and short- and long-chain fatty acids have been tested with respect to their potential to influence epigenetic mechanisms. Considering that a complex mixture of intermediary and microbial metabolites is present in human circulation, a more systematic interdisciplinary investigation of nutri-epigenetic activities and their impact on human health is called for.This article is part of a discussion meeting issue 'Frontiers in epigenetic chemical biology'.
Keywords: diet; epigenomics; gut microbiota; human health; metabolism.
© 2018 The Authors.
Conflict of interest statement
I declare I have no competing interests.
Figures
Similar articles
-
Gut Microbiota as Important Mediator Between Diet and DNA Methylation and Histone Modifications in the Host.Nutrients. 2020 Feb 25;12(3):597. doi: 10.3390/nu12030597. Nutrients. 2020. PMID: 32106534 Free PMC article. Review.
-
Dietary metabolites derived from gut microbiota: critical modulators of epigenetic changes in mammals.Nutr Rev. 2017 May 1;75(5):374-389. doi: 10.1093/nutrit/nux001. Nutr Rev. 2017. PMID: 28444216 Review.
-
More than just a gut instinct-the potential interplay between a baby's nutrition, its gut microbiome, and the epigenome.Am J Physiol Regul Integr Comp Physiol. 2013 Jun 15;304(12):R1065-9. doi: 10.1152/ajpregu.00551.2012. Epub 2013 Apr 17. Am J Physiol Regul Integr Comp Physiol. 2013. PMID: 23594611
-
The interplay between diet, gut microbes, and host epigenetics in health and disease.J Nutr Biochem. 2021 Sep;95:108631. doi: 10.1016/j.jnutbio.2021.108631. Epub 2021 Mar 28. J Nutr Biochem. 2021. PMID: 33789148 Free PMC article. Review.
-
Integration of microbiome and epigenome to decipher the pathogenesis of autoimmune diseases.J Autoimmun. 2017 Sep;83:31-42. doi: 10.1016/j.jaut.2017.03.009. Epub 2017 Mar 23. J Autoimmun. 2017. PMID: 28342734 Review.
Cited by
-
What We Know So Far about the Metabolite-Mediated Microbiota-Intestinal Immunity Dialogue and How to Hear the Sound of This Crosstalk.Metabolites. 2021 Jun 21;11(6):406. doi: 10.3390/metabo11060406. Metabolites. 2021. PMID: 34205653 Free PMC article. Review.
-
Live yeast in juvenile diet induces species-specific effects on Drosophila adult behaviour and fitness.Sci Rep. 2019 Jun 20;9(1):8873. doi: 10.1038/s41598-019-45140-z. Sci Rep. 2019. PMID: 31222019 Free PMC article.
-
Microalgae, Seaweeds and Aquatic Bacteria, Archaea, and Yeasts: Sources of Carotenoids with Potential Antioxidant and Anti-Inflammatory Health-Promoting Actions in the Sustainability Era.Mar Drugs. 2023 Jun 1;21(6):340. doi: 10.3390/md21060340. Mar Drugs. 2023. PMID: 37367666 Free PMC article. Review.
-
Feed Composition Differences Resulting from Organic and Conventional Farming Practices Affect Physiological Parameters in Wistar Rats-Results from a Factorial, Two-Generation Dietary Intervention Trial.Nutrients. 2021 Jan 26;13(2):377. doi: 10.3390/nu13020377. Nutrients. 2021. PMID: 33530419 Free PMC article.
-
The interaction between gut microbiome and nutrients on development of human disease through epigenetic mechanisms.Genomics Inform. 2019 Sep;17(3):e24. doi: 10.5808/GI.2019.17.3.e24. Epub 2019 Sep 26. Genomics Inform. 2019. PMID: 31610620 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous