A Node-Expressed Transporter OsCCX2 Is Involved in Grain Cadmium Accumulation of Rice
- PMID: 29696032
- PMCID: PMC5904359
- DOI: 10.3389/fpls.2018.00476
A Node-Expressed Transporter OsCCX2 Is Involved in Grain Cadmium Accumulation of Rice
Abstract
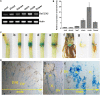
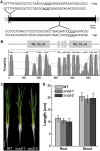
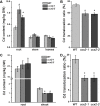
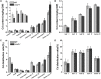
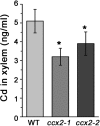
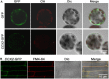
Excessive cadmium (Cd) accumulation in grains of rice (Oryza sativa L.) is a risk to food security. The transporters in the nodes of rice are involved in the distribution of mineral elements including toxic elements to different tissues such as grains. However, the mechanism of Cd accumulation in grains is largely unknown. Here, we report a node-expressed transporter gene, OsCCX2, a putative cation/calcium (Ca) exchanger, mediating Cd accumulation in the grains of rice. Knockout of OsCCX2 caused a remarkable reduction of Cd content in the grains. Further study showed that disruption of this gene led to a reduced root-to-shoot translocation ratio of Cd. Moreover, Cd distribution was also disturbed in different levels of internode and leaf. OsCCX2 is localized to plasma membrane, and OsCCX2 is mainly expressed in xylem region of vascular tissues at the nodes. OsCCX2 might function as an efflux transporter, responsible for Cd loading into xylem vessels. Therefore, our finding revealed a novel Cd transporter involved in grain Cd accumulation, possibly via a Ca transport pathway in the nodes of rice.
Keywords: Oryza sativa; cadmium accumulation; distribution; translocation; transporter.
Figures


Similar articles
-
Research Advances in Cadmium Uptake, Transport and Resistance in Rice (Oryza sativa L.).Cells. 2022 Feb 6;11(3):569. doi: 10.3390/cells11030569. Cells. 2022. PMID: 35159378 Free PMC article. Review.
-
Route and Regulation of Zinc, Cadmium, and Iron Transport in Rice Plants (Oryza sativa L.) during Vegetative Growth and Grain Filling: Metal Transporters, Metal Speciation, Grain Cd Reduction and Zn and Fe Biofortification.Int J Mol Sci. 2015 Aug 13;16(8):19111-29. doi: 10.3390/ijms160819111. Int J Mol Sci. 2015. PMID: 26287170 Free PMC article. Review.
-
Low-affinity cation transporter (OsLCT1) regulates cadmium transport into rice grains.Proc Natl Acad Sci U S A. 2011 Dec 27;108(52):20959-64. doi: 10.1073/pnas.1116531109. Epub 2011 Dec 12. Proc Natl Acad Sci U S A. 2011. PMID: 22160725 Free PMC article.
-
Root-to-shoot Cd translocation via the xylem is the major process determining shoot and grain cadmium accumulation in rice.J Exp Bot. 2009;60(9):2677-88. doi: 10.1093/jxb/erp119. Epub 2009 Apr 28. J Exp Bot. 2009. PMID: 19401409 Free PMC article.
-
Effects and mechanisms of foliar application of silicon and selenium composite sols on diminishing cadmium and lead translocation and affiliated physiological and biochemical responses in hybrid rice (Oryza sativa L.) exposed to cadmium and lead.Chemosphere. 2020 Jul;251:126347. doi: 10.1016/j.chemosphere.2020.126347. Epub 2020 Feb 27. Chemosphere. 2020. PMID: 32169700
Cited by
-
(De)Activation (Ir)Reversibly or Degradation: Dynamics of Post-Translational Protein Modifications in Plants.Life (Basel). 2022 Feb 21;12(2):324. doi: 10.3390/life12020324. Life (Basel). 2022. PMID: 35207610 Free PMC article. Review.
-
Research Advances in Cadmium Uptake, Transport and Resistance in Rice (Oryza sativa L.).Cells. 2022 Feb 6;11(3):569. doi: 10.3390/cells11030569. Cells. 2022. PMID: 35159378 Free PMC article. Review.
-
Advances in molecular mechanisms underlying cadmium uptake and translocation in rice.Front Plant Sci. 2022 Sep 20;13:1003953. doi: 10.3389/fpls.2022.1003953. eCollection 2022. Front Plant Sci. 2022. PMID: 36204081 Free PMC article. Review.
-
Advances in the Uptake and Transport Mechanisms and QTLs Mapping of Cadmium in Rice.Int J Mol Sci. 2019 Jul 11;20(14):3417. doi: 10.3390/ijms20143417. Int J Mol Sci. 2019. PMID: 31336794 Free PMC article. Review.
-
Transcriptional Regulatory Network of Plant Cadmium Stress Response.Int J Mol Sci. 2023 Feb 22;24(5):4378. doi: 10.3390/ijms24054378. Int J Mol Sci. 2023. PMID: 36901809 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

