The 3' mRNA I-shaped structure of maize necrotic streak virus binds to eukaryotic translation factors for eIF4F-mediated translation initiation
- PMID: 29700118
- PMCID: PMC6005423
- DOI: 10.1074/jbc.RA118.003377
The 3' mRNA I-shaped structure of maize necrotic streak virus binds to eukaryotic translation factors for eIF4F-mediated translation initiation
Abstract
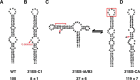
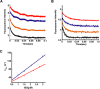
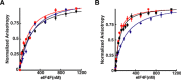
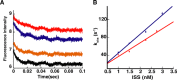
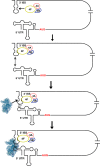
Unlike the mRNAs of their eukaryotic hosts, many RNAs of viruses lack a 5' m7GpppN cap and the 3' polyadenosine tail, and yet they are translated efficiently. Plant RNA viruses, in particular, have complex structures within their mRNA UTRs that allow them to bypass some cellular translation control steps. In the 3' UTR of maize necrotic streak virus (MNeSV), an I-shaped RNA structure (ISS) has been shown to bind eukaryotic initiation factor (eIF)4F and to mediate viral translation initiation. A 5'-3' RNA "kissing-loop" interaction is required for optimal translation. However, the details of how the 3' ISS mediates translation initiation are not well understood. Here, we studied the binding of the 3' ISS with eIFs. The eIF4A-eIF4B complex was found to increase binding affinity of eIF4F with the 3' ISS by 4-fold (from KD = 173 ± 34 nm to KD = 48 ± 11 nm). Pre-steady-state analysis indicated that the eIF4A-eIF4B complex increased the RNA association rate and decreased the dissociation rate in an ATP-independent manner. Furthermore, our findings suggest that eIF4F could promote binding of the 3' ISS with the MNeSV 5'UTR, enhancing the long-distance kissing-loop interaction. However, the association of the 5'UTR with the 3' ISS-eIF4F complex did not increase 40S ribosomal subunit binding affinity. These quantitative results suggest a stepwise model in which the first committed step is eIF4F binding to the 3' ISS, followed by an interaction with the 5'UTR and subsequent 40S ribosomal subunit binding.
Keywords: I-shaped RNA structure; RNA UTR; RNA structure; RNA virus; RNA–protein interaction; eukaryotic translation initiation; kissing loop; maize necrotic streak virus; plant virus; viral protein translation.
© 2018 Liu and Goss.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures



Similar articles
-
Eukaryotic initiation factor (eIF) 3 mediates Barley Yellow Dwarf Viral mRNA 3'-5' UTR interactions and 40S ribosomal subunit binding to facilitate cap-independent translation.Nucleic Acids Res. 2019 Jul 9;47(12):6225-6235. doi: 10.1093/nar/gkz448. Nucleic Acids Res. 2019. PMID: 31114905 Free PMC article.
-
Recruitment of the 40S ribosome subunit to the 3'-untranslated region (UTR) of a viral mRNA, via the eIF4 complex, facilitates cap-independent translation.J Biol Chem. 2015 May 1;290(18):11268-81. doi: 10.1074/jbc.M115.645002. Epub 2015 Mar 19. J Biol Chem. 2015. PMID: 25792742 Free PMC article.
-
Evidence for variation in the optimal translation initiation complex: plant eIF4B, eIF4F, and eIF(iso)4F differentially promote translation of mRNAs.Plant Physiol. 2009 Aug;150(4):1844-54. doi: 10.1104/pp.109.138438. Epub 2009 Jun 3. Plant Physiol. 2009. PMID: 19493973 Free PMC article.
-
eIF4F: a retrospective.J Biol Chem. 2015 Oct 2;290(40):24091-9. doi: 10.1074/jbc.R115.675280. Epub 2015 Aug 31. J Biol Chem. 2015. PMID: 26324716 Free PMC article. Review.
-
Hepatitis C Virus Translation Regulation.Int J Mol Sci. 2020 Mar 27;21(7):2328. doi: 10.3390/ijms21072328. Int J Mol Sci. 2020. PMID: 32230899 Free PMC article. Review.
Cited by
-
Opium Poppy Mosaic Virus Has an Xrn-Resistant, Translated Subgenomic RNA and a BTE 3' CITE.J Virol. 2021 Apr 12;95(9):e02109-20. doi: 10.1128/JVI.02109-20. Print 2021 Apr 12. J Virol. 2021. PMID: 33597210 Free PMC article.
-
Mechanistic differences in eukaryotic initiation factor requirements for eIF4GI-driven cap-independent translation of structured mRNAs.J Biol Chem. 2024 Nov;300(11):107866. doi: 10.1016/j.jbc.2024.107866. Epub 2024 Oct 9. J Biol Chem. 2024. PMID: 39384039 Free PMC article.
-
Identification of Novel 5' and 3' Translation Enhancers in Umbravirus-Like Coat Protein-Deficient RNA Replicons.J Virol. 2022 Apr 13;96(7):e0173621. doi: 10.1128/jvi.01736-21. Epub 2022 Mar 17. J Virol. 2022. PMID: 35297668 Free PMC article.
-
Translational gene regulation in plants: A green new deal.Wiley Interdiscip Rev RNA. 2020 Nov;11(6):e1597. doi: 10.1002/wrna.1597. Epub 2020 May 4. Wiley Interdiscip Rev RNA. 2020. PMID: 32367681 Free PMC article. Review.
-
eIF4E1 Regulates Arabidopsis Embryo Development and Root Growth by Interacting With RopGEF7.Front Plant Sci. 2022 Jun 30;13:938476. doi: 10.3389/fpls.2022.938476. eCollection 2022. Front Plant Sci. 2022. PMID: 35845661 Free PMC article.
References
-
- De Stradis, A., Redinbaugh, M. G., Abt, J. J., and Martelli, G. P. (2005) Ultrastructure of maize necrotic streak virus infections. J. Plant Pathol. 87, 213–221
-
- Stuart, G., Moffett, K., and Bozarth, R. F. (2004) A whole genome perspective on the phylogeny of the plant virus family Tombusviridae. Arch. Virol. 149, 1595–1610 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

