Genome-centric metatranscriptomes and ecological roles of the active microbial populations during cellulosic biomass anaerobic digestion
- PMID: 29713376
- PMCID: PMC5911951
- DOI: 10.1186/s13068-018-1121-0
Genome-centric metatranscriptomes and ecological roles of the active microbial populations during cellulosic biomass anaerobic digestion
Abstract
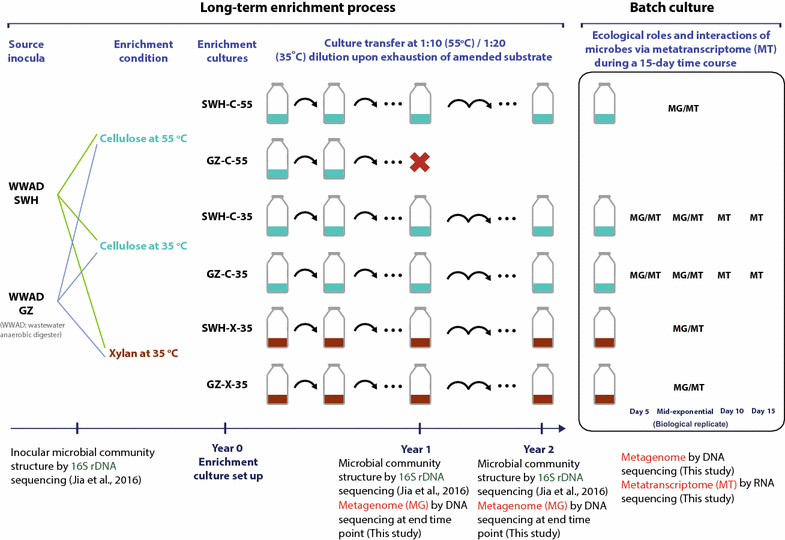
Background: Although anaerobic digestion for biogas production is used worldwide in treatment processes to recover energy from carbon-rich waste such as cellulosic biomass, the activities and interactions among the microbial populations that perform anaerobic digestion deserve further investigations, especially at the population genome level. To understand the cellulosic biomass-degrading potentials in two full-scale digesters, this study examined five methanogenic enrichment cultures derived from the digesters that anaerobically digested cellulose or xylan for more than 2 years under 35 or 55 °C conditions.
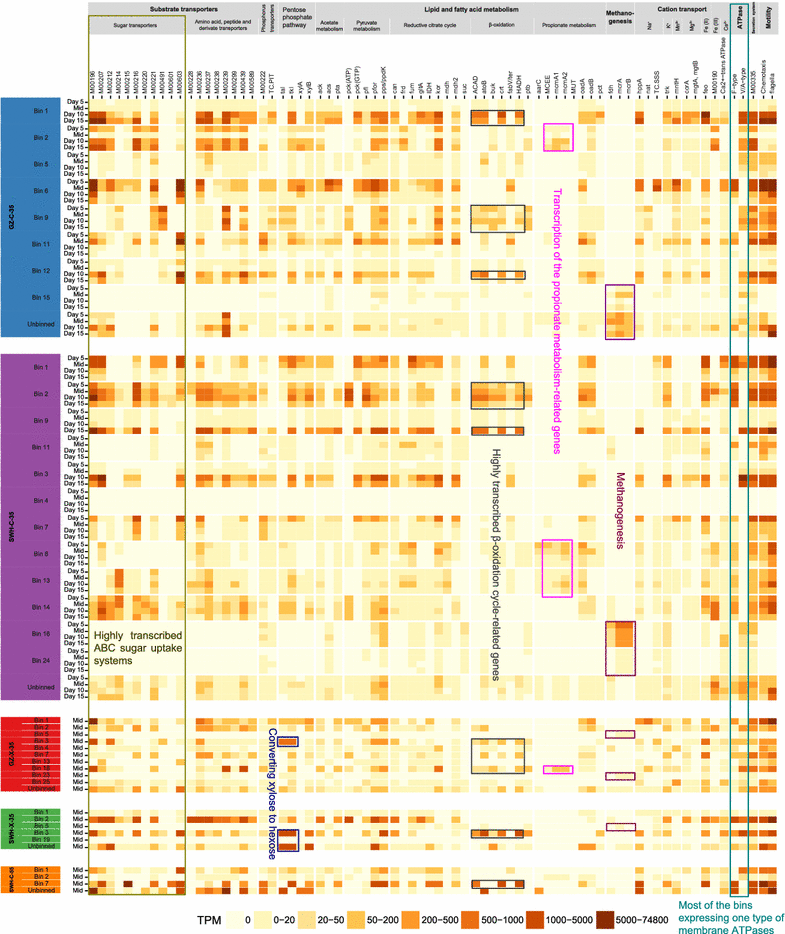
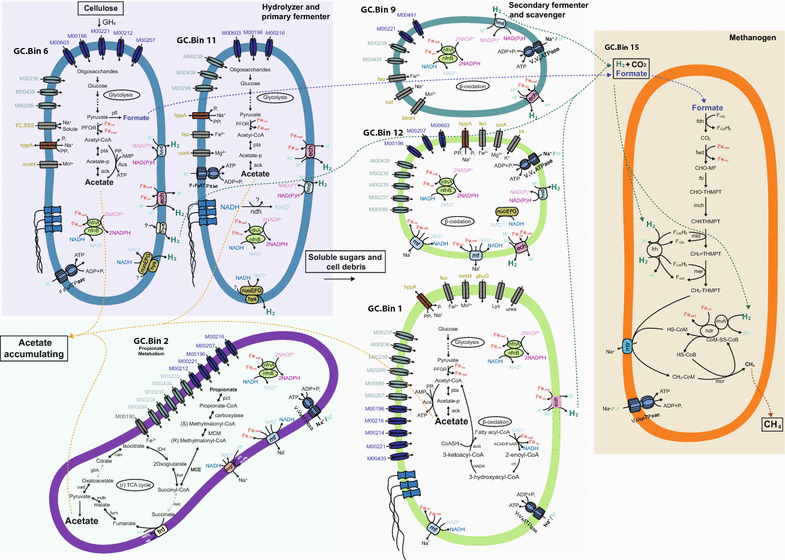
Results: Metagenomics and metatranscriptomics were used to capture the active microbial populations in each enrichment culture and reconstruct their meta-metabolic network and ecological roles. 107 population genomes were reconstructed from the five enrichment cultures using a differential coverage binning approach, of which only a subset was highly transcribed in the metatranscriptomes. Phylogenetic and functional convergence of communities by enrichment condition and phase of fermentation was observed for the highly transcribed populations in the metatranscriptomes. In the 35 °C cultures grown on cellulose, Clostridium cellulolyticum-related and Ruminococcus-related bacteria were identified as major hydrolyzers and primary fermenters in the early growth phase, while Clostridium leptum-related bacteria were major secondary fermenters and potential fatty acid scavengers in the late growth phase. While the meta-metabolism and trophic roles of the cultures were similar, the bacterial populations performing each function were distinct between the enrichment conditions.
Conclusions: Overall, a population genome-centric view of the meta-metabolism and functional roles of key active players in anaerobic digestion of cellulosic biomass was obtained. This study represents a major step forward towards understanding the microbial functions and interactions at population genome level during the microbial conversion of lignocellulosic biomass to methane. The knowledge of this study can facilitate development of potential biomarkers and rational design of the microbiome in anaerobic digesters.
Keywords: Anaerobic digestion; Ecological roles; Lignocellulosic biomass; Metagenomics; Metatranscriptomics.
Figures


Similar articles
-
Upflow anaerobic sludge blanket reactor--a review.Indian J Environ Health. 2001 Apr;43(2):1-82. Indian J Environ Health. 2001. PMID: 12397675 Review.
-
Elucidating Syntrophic Butyrate-Degrading Populations in Anaerobic Digesters Using Stable-Isotope-Informed Genome-Resolved Metagenomics.mSystems. 2019 Aug 6;4(4):e00159-19. doi: 10.1128/mSystems.00159-19. mSystems. 2019. PMID: 31387934 Free PMC article.
-
Genome-centric resolution of microbial diversity, metabolism and interactions in anaerobic digestion.Environ Microbiol. 2016 Sep;18(9):3144-58. doi: 10.1111/1462-2920.13382. Epub 2016 Jul 22. Environ Microbiol. 2016. PMID: 27317862
-
Long-Term Enrichment on Cellulose or Xylan Causes Functional and Taxonomic Convergence of Microbial Communities from Anaerobic Digesters.Appl Environ Microbiol. 2015 Dec 28;82(5):1519-1529. doi: 10.1128/AEM.03360-15. Appl Environ Microbiol. 2015. PMID: 26712547 Free PMC article.
-
Application of next-generation sequencing methods for microbial monitoring of anaerobic digestion of lignocellulosic biomass.Appl Microbiol Biotechnol. 2017 Sep;101(18):6849-6864. doi: 10.1007/s00253-017-8438-7. Epub 2017 Aug 4. Appl Microbiol Biotechnol. 2017. PMID: 28779289 Review.
Cited by
-
High biodiversity in a benzene-degrading nitrate-reducing culture is sustained by a few primary consumers.Commun Biol. 2021 May 5;4(1):530. doi: 10.1038/s42003-021-01948-y. Commun Biol. 2021. PMID: 33953314 Free PMC article.
-
A microbial gene catalog of anaerobic digestion from full-scale biogas plants.Gigascience. 2021 Jan 27;10(1):giaa164. doi: 10.1093/gigascience/giaa164. Gigascience. 2021. PMID: 33506264 Free PMC article.
-
Molecular Microbial Community Analysis as an Analysis Tool for Optimal Biogas Production.Microorganisms. 2021 May 28;9(6):1162. doi: 10.3390/microorganisms9061162. Microorganisms. 2021. PMID: 34071282 Free PMC article. Review.
-
A unified compendium of prokaryotic and viral genomes from over 300 anaerobic digestion microbiomes.Environ Microbiome. 2024 Jan 2;19(1):1. doi: 10.1186/s40793-023-00545-2. Environ Microbiome. 2024. PMID: 38167520 Free PMC article.
-
Metatranscriptomics-guided genome-scale metabolic reconstruction reveals the carbon flux and trophic interaction in methanogenic communities.Microbiome. 2024 Jul 5;12(1):121. doi: 10.1186/s40168-024-01830-z. Microbiome. 2024. PMID: 38970122 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

