Endocardial Hippo signaling regulates myocardial growth and cardiogenesis
- PMID: 29727635
- PMCID: PMC5989000
- DOI: 10.1016/j.ydbio.2018.04.026
Endocardial Hippo signaling regulates myocardial growth and cardiogenesis
Abstract
The Hippo signaling pathway has been implicated in control of cell and organ size, proliferation, and endothelial-mesenchymal transformation. This pathway impacts upon two partially redundant transcription cofactors, Yap and Taz, that interact with other factors, including members of the Tead family, to affect expression of downstream genes. Yap and Taz have been shown to regulate, in a cell-autonomous manner, myocardial proliferation, myocardial hypertrophy, regenerative potential, and overall size of the heart. Here, we show that Yap and Taz also play an instructive, non-cell-autonomous role in the endocardium of the developing heart to regulate myocardial growth through release of the paracrine factor, neuregulin. Without endocardial Yap and Taz, myocardial growth is impaired causing early post-natal lethality. Thus, the Hippo signaling pathway regulates cell size via both cell-autonomous and non-cell-autonomous mechanisms. Furthermore, these data suggest that Hippo may regulate organ size via a sensing and paracrine function in endothelial cells.
Copyright © 2018 The Authors. Published by Elsevier Inc. All rights reserved.
Figures



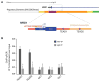


References
-
- Bersell K, Arab S, Haring B, Kuhn B. Neuregulin1/ErbB4 signaling induces cardiomyocyte proliferation and repair of heart injury. Cell. 2009;138:257–270. - PubMed
-
- Brutsaert DL. Cardiac endothelial-myocardial signaling: its role in cardiac growth, contractile performance, and rhythmicity. Physiol Rev. 2003;83:59–115. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

