Ensemble outlier detection and gene selection in triple-negative breast cancer data
- PMID: 29728051
- PMCID: PMC5936001
- DOI: 10.1186/s12859-018-2149-7
Ensemble outlier detection and gene selection in triple-negative breast cancer data
Abstract
Background: Learning accurate models from 'omics data is bringing many challenges due to their inherent high-dimensionality, e.g. the number of gene expression variables, and comparatively lower sample sizes, which leads to ill-posed inverse problems. Furthermore, the presence of outliers, either experimental errors or interesting abnormal clinical cases, may severely hamper a correct classification of patients and the identification of reliable biomarkers for a particular disease. We propose to address this problem through an ensemble classification setting based on distinct feature selection and modeling strategies, including logistic regression with elastic net regularization, Sparse Partial Least Squares - Discriminant Analysis (SPLS-DA) and Sparse Generalized PLS (SGPLS), coupled with an evaluation of the individuals' outlierness based on the Cook's distance. The consensus is achieved with the Rank Product statistics corrected for multiple testing, which gives a final list of sorted observations by their outlierness level.
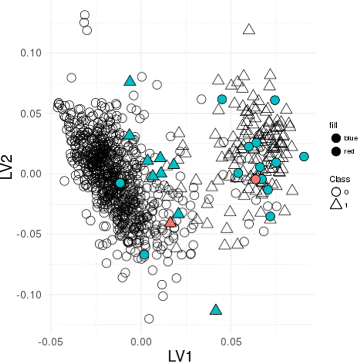
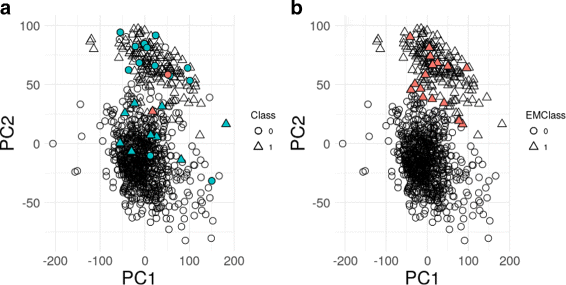
Results: We applied this strategy for the classification of Triple-Negative Breast Cancer (TNBC) RNA-Seq and clinical data from the Cancer Genome Atlas (TCGA). The detected 24 outliers were identified as putative mislabeled samples, corresponding to individuals with discrepant clinical labels for the HER2 receptor, but also individuals with abnormal expression values of ER, PR and HER2, contradictory with the corresponding clinical labels, which may invalidate the initial TNBC label. Moreover, the model consensus approach leads to the selection of a set of genes that may be linked to the disease. These results are robust to a resampling approach, either by selecting a subset of patients or a subset of genes, with a significant overlap of the outlier patients identified.
Conclusions: The proposed ensemble outlier detection approach constitutes a robust procedure to identify abnormal cases and consensus covariates, which may improve biomarker selection for precision medicine applications. The method can also be easily extended to other regression models and datasets.
Keywords: Ensemble modeling; High-dimensionality; Outlier detection; Rank Product test; Triple-negative breast cancer.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Robust identification of target genes and outliers in triple-negative breast cancer data.Stat Methods Med Res. 2019 Oct-Nov;28(10-11):3042-3056. doi: 10.1177/0962280218794722. Epub 2018 Aug 27. Stat Methods Med Res. 2019. PMID: 30146936 Free PMC article.
-
Comparison of methods for the detection of outliers and associated biomarkers in mislabeled omics data.BMC Bioinformatics. 2020 Aug 14;21(1):357. doi: 10.1186/s12859-020-03653-9. BMC Bioinformatics. 2020. PMID: 32795265 Free PMC article.
-
ROSIE: RObust Sparse ensemble for outlIEr detection and gene selection in cancer omics data.Stat Methods Med Res. 2022 May;31(5):947-958. doi: 10.1177/09622802211072456. Epub 2022 Jan 24. Stat Methods Med Res. 2022. PMID: 35072570 Free PMC article.
-
Long noncoding RNAs (lncRNAs) in triple negative breast cancer.J Cell Physiol. 2017 Dec;232(12):3226-3233. doi: 10.1002/jcp.25830. Epub 2017 Feb 21. J Cell Physiol. 2017. PMID: 28138992 Review.
-
MicroRNAs-mediated cell fate in triple negative breast cancers.Cancer Lett. 2015 May 28;361(1):8-12. doi: 10.1016/j.canlet.2015.02.048. Epub 2015 Mar 3. Cancer Lett. 2015. PMID: 25748387 Review.
Cited by
-
TCox: Correlation-Based Regularization Applied to Colorectal Cancer Survival Data.Biomedicines. 2020 Nov 10;8(11):488. doi: 10.3390/biomedicines8110488. Biomedicines. 2020. PMID: 33182598 Free PMC article.
-
DNA Methylation and Breast Cancer Risk: An Epigenome-Wide Study of Normal Breast Tissue and Blood.Cancers (Basel). 2020 Oct 23;12(11):3088. doi: 10.3390/cancers12113088. Cancers (Basel). 2020. PMID: 33113958 Free PMC article.
-
Sialyl LewisX/A and Cytokeratin Crosstalk in Triple Negative Breast Cancer.Cancers (Basel). 2023 Jan 25;15(3):731. doi: 10.3390/cancers15030731. Cancers (Basel). 2023. PMID: 36765690 Free PMC article.
-
An Efficient Algorithm for the Detection of Outliers in Mislabeled Omics Data.Comput Math Methods Med. 2021 Dec 22;2021:9436582. doi: 10.1155/2021/9436582. eCollection 2021. Comput Math Methods Med. 2021. PMID: 34976114 Free PMC article.
-
Robust identification of target genes and outliers in triple-negative breast cancer data.Stat Methods Med Res. 2019 Oct-Nov;28(10-11):3042-3056. doi: 10.1177/0962280218794722. Epub 2018 Aug 27. Stat Methods Med Res. 2019. PMID: 30146936 Free PMC article.
References
-
- Basu B, Basu S. Correlating and combining genomic and proteomic assessment with in vivo molecular functional imaging: Will this be the future roadmap for personalized cancer management? Nat Med. 2016;31(3):75–84. - PubMed
-
- Aggarwal CC. Outlier ensembles [position paper] ACM SIGKDD Explor. 2012;14(49-58):2.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

