Identification of the Intragenomic Promoter Controlling Hepatitis E Virus Subgenomic RNA Transcription
- PMID: 29739903
- PMCID: PMC5941075
- DOI: 10.1128/mBio.00769-18
Identification of the Intragenomic Promoter Controlling Hepatitis E Virus Subgenomic RNA Transcription
Abstract
Approximately 20 million hepatitis E virus (HEV) infections occur annually in both developing and industrialized countries. Most infections are self-limiting, but they can lead to chronic infections and cirrhosis in immunocompromised patients, and death in pregnant women. The mechanisms of HEV replication remain incompletely understood due to scarcity of adequate experimental platforms. HEV undergoes asymmetric genome replication, but it produces an additional subgenomic (SG) RNA encoding the viral capsid and a viroporin in partially overlapping open reading frames. Using a novel transcomplementation system, we mapped the intragenomic subgenomic promoter regulating SG RNA synthesis. This cis-acting element is highly conserved across all eight HEV genotypes, and when the element is mutated, it abrogates particle assembly and release. Our work defines previously unappreciated viral regulatory elements and provides the first in-depth view of the intracellular genome dynamics of this emerging human pathogen.IMPORTANCE HEV is an emerging pathogen causing severe liver disease. The genetic information of HEV is encoded in RNA. The genomic RNA is initially copied into a complementary, antigenomic RNA that is a template for synthesis of more genomic RNA and for so-called subgenomic RNA. In this study, we identified the precise region within the HEV genome at which the synthesis of the subgenomic RNA is initiated. The nucleotides within this region are conserved across genetically distinct variants of HEV, highlighting the general importance of this segment for the virus. To identify this regulatory element, we developed a new experimental system that is a powerful tool with broad utility to mechanistically dissect many other poorly understood functional elements of HEV.
Keywords: hepatitis E; hepatitis E virus; viral hepatitis; viral replication.
Copyright © 2018 Ding et al.
Figures







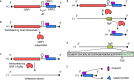
References
-
- Koonin EV, Gorbalenya AE, Purdy MA, Rozanov MN, Reyes GR, Bradley DW. 1992. Computer-assisted assignment of functional domains in the nonstructural polyprotein of hepatitis E virus: delineation of an additional group of positive-strand RNA plant and animal viruses. Proc Natl Acad Sci U S A 89:8259–8263. doi: 10.1073/pnas.89.17.8259. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
