Mechanism of Action of Mycobacterium tuberculosis Gyrase Inhibitors: A Novel Class of Gyrase Poisons
- PMID: 29746087
- PMCID: PMC6309371
- DOI: 10.1021/acsinfecdis.8b00035
Mechanism of Action of Mycobacterium tuberculosis Gyrase Inhibitors: A Novel Class of Gyrase Poisons
Abstract
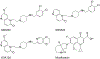
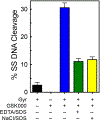
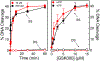
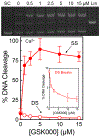
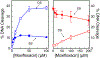
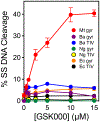
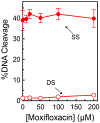
Tuberculosis is one of the leading causes of morbidity worldwide, and the incidences of drug resistance and intolerance are prevalent. Thus, there is a desperate need for the development of new antitubercular drugs. Mycobacterium tuberculosis gyrase inhibitors (MGIs) are napthyridone/aminopiperidine-based drugs that display activity against M. tuberculosis cells and tuberculosis in mouse models [Blanco, D., et al. (2015) Antimicrob. Agents Chemother. 59, 1868-1875]. Genetic and mutagenesis studies suggest that gyrase, which is the target for fluoroquinolone antibacterials, is also the target for MGIs. However, little is known regarding the interaction of these drugs with the bacterial type II enzyme. Therefore, we examined the effects of two MGIs, GSK000 and GSK325, on M. tuberculosis gyrase. MGIs greatly enhanced DNA cleavage mediated by the bacterial enzyme. In contrast to fluoroquinolones (which induce primarily double-stranded breaks), MGIs induced only single-stranded DNA breaks under a variety of conditions. MGIs work by stabilizing covalent gyrase-cleaved DNA complexes and appear to suppress the ability of the enzyme to induce double-stranded breaks. The drugs displayed little activity against type II topoisomerases from several other bacterial species, suggesting that these drugs display specificity for M. tuberculosis gyrase. Furthermore, MGIs maintained activity against M. tuberuclosis gyrase enzymes that contained the three most common fluoroquinolone resistance mutations seen in the clinic and displayed no activity against human topoisomerase IIα. These findings suggest that MGIs have potential as antitubercular drugs, especially in the case of fluoroquinolone-resistant disease.
Keywords: Mycobacterium tuberculosis gyrase inhibitors; fluoroquinolones; gyrase; single-stranded DNA cleavage; tuberculosis.
Conflict of interest statement
The authors declare no competing financial interests.
Figures






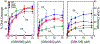

Similar articles
-
Bimodal Actions of a Naphthyridone/Aminopiperidine-Based Antibacterial That Targets Gyrase and Topoisomerase IV.Biochemistry. 2019 Nov 5;58(44):4447-4455. doi: 10.1021/acs.biochem.9b00805. Epub 2019 Oct 28. Biochemistry. 2019. PMID: 31617352 Free PMC article.
-
A Series of Spiropyrimidinetriones that Enhances DNA Cleavage Mediated by Mycobacterium tuberculosis Gyrase.ACS Infect Dis. 2023 Mar 10;9(3):706-715. doi: 10.1021/acsinfecdis.3c00012. Epub 2023 Feb 20. ACS Infect Dis. 2023. PMID: 36802491 Free PMC article.
-
Targeting DNA Gyrase to Combat Mycobacterium tuberculosis: An Update.Curr Top Med Chem. 2019;19(8):579-593. doi: 10.2174/1568026619666190304130218. Curr Top Med Chem. 2019. PMID: 30834837 Review.
-
Novel N-linked aminopiperidine-based gyrase inhibitors with improved hERG and in vivo efficacy against Mycobacterium tuberculosis.J Med Chem. 2014 Jun 12;57(11):4889-905. doi: 10.1021/jm500432n. Epub 2014 May 23. J Med Chem. 2014. PMID: 24809953
-
Gyrase and Topoisomerase IV: Recycling Old Targets for New Antibacterials to Combat Fluoroquinolone Resistance.ACS Infect Dis. 2024 Apr 12;10(4):1097-1115. doi: 10.1021/acsinfecdis.4c00128. Epub 2024 Apr 2. ACS Infect Dis. 2024. PMID: 38564341 Free PMC article. Review.
Cited by
-
Spiropyrimidinetriones: a Class of DNA Gyrase Inhibitors with Activity against Mycobacterium tuberculosis and without Cross-Resistance to Fluoroquinolones.Antimicrob Agents Chemother. 2022 Apr 19;66(4):e0219221. doi: 10.1128/aac.02192-21. Epub 2022 Mar 10. Antimicrob Agents Chemother. 2022. PMID: 35266826 Free PMC article.
-
Dioxane-Linked Amide Derivatives as Novel Bacterial Topoisomerase Inhibitors against Gram-Positive Staphylococcus aureus.ACS Med Chem Lett. 2020 Oct 19;11(12):2446-2454. doi: 10.1021/acsmedchemlett.0c00428. eCollection 2020 Dec 10. ACS Med Chem Lett. 2020. PMID: 33335666 Free PMC article.
-
Actions of a Novel Bacterial Topoisomerase Inhibitor against Neisseria gonorrhoeae Gyrase and Topoisomerase IV: Enhancement of Double-Stranded DNA Breaks.Int J Mol Sci. 2023 Jul 28;24(15):12107. doi: 10.3390/ijms241512107. Int J Mol Sci. 2023. PMID: 37569485 Free PMC article.
-
A Mycobacterium tuberculosis NBTI DNA Gyrase Inhibitor Is Active against Mycobacterium abscessus.Antimicrob Agents Chemother. 2021 Nov 17;65(12):e0151421. doi: 10.1128/AAC.01514-21. Epub 2021 Oct 4. Antimicrob Agents Chemother. 2021. PMID: 34606340 Free PMC article.
-
Bimodal Actions of a Naphthyridone/Aminopiperidine-Based Antibacterial That Targets Gyrase and Topoisomerase IV.Biochemistry. 2019 Nov 5;58(44):4447-4455. doi: 10.1021/acs.biochem.9b00805. Epub 2019 Oct 28. Biochemistry. 2019. PMID: 31617352 Free PMC article.
References
-
- Namdar R; Lauzardo M; Peloquin CA, Tuberculosis In Pharmacotherapy : a pathophysiologic approach, 9th edition ed.; DiPiro JT, Ed. McGraw-Hill: New York, 2014.
-
- WHO, Global tuberculosis report 2016. 2016.
-
- WHO Tuberculosis Fact Sheet http://www.who.int/mediacentre/factsheets/fs104/en/.
-
- WHO, Treatment of tuberculosis: guidelines. 4th ed.; WHO Press: Geneva, 2010.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

