Bifunctional cross-linking approaches for mass spectrometry-based investigation of nucleic acids and protein-nucleic acid assemblies
- PMID: 29753003
- PMCID: PMC7034459
- DOI: 10.1016/j.ymeth.2018.05.001
Bifunctional cross-linking approaches for mass spectrometry-based investigation of nucleic acids and protein-nucleic acid assemblies
Abstract
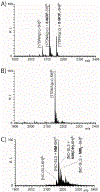
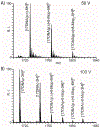
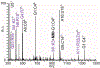
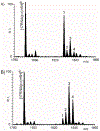
With the goal of expanding the very limited toolkit of cross-linking agents available for nucleic acids and their protein complexes, we evaluated the merits of a wide range of bifunctional agents that may be capable of reacting with the functional groups characteristic of these types of biopolymers. The survey specifically focused on the ability of test reagents to produce desirable inter-molecular conjugates, which could reveal the identity of interacting components and the position of mutual contacts, while also considering a series of practical criteria for their utilization as viable nucleic acid probes. The survey employed models consisting of DNA, RNA, and corresponding protein complexes to mimic as close as possible typical applications. Denaturing polyacrylamide gel electrophoresis (PAGE) and mass spectrometric (MS) analyses were implemented in concert to monitor the formation of the desired conjugates. In particular, the former was used as a rapid and inexpensive tool for the efficient evaluation of cross-linker activity under a broad range of experimental conditions. The latter was applied after preliminary rounds of reaction optimization to enable full-fledged product characterization and, more significantly, differentiation between mono-functional and intra- versus inter-molecular conjugates. This information provided the feedback necessary to further optimize reaction conditions and explain possible outcomes. Among the reagents tested in the study, platinum complexes and nitrogen mustards manifested the most favorable characteristics for practical cross-linking applications, whereas other compounds provided inferior yields, or produced rather unstable conjugates that did not survive the selected analytical conditions. The observed outcomes will help guide the selection of the most appropriate cross-linking reagent for a specific task, whereas the experimental conditions described here will provide an excellent starting point for approaching these types of applications. As a whole, the results of the survey clearly emphasize that finding a universal reagent, which may afford excellent performance with all types of nucleic acid substrates, will require extending the exploration beyond the traditional chemistries employed to modify the constitutive functional groups of these vital biopolymers.
Copyright © 2018. Published by Elsevier Inc.
Figures



Similar articles
-
Top-down characterization of nucleic acids modified by structural probes using high-resolution tandem mass spectrometry and automated data interpretation.Anal Chem. 2004 May 1;76(9):2438-45. doi: 10.1021/ac0355045. Anal Chem. 2004. PMID: 15117181
-
Analysis of protein-nucleic acid interactions by photochemical cross-linking and mass spectrometry.Mass Spectrom Rev. 2002 May-Jun;21(3):163-82. doi: 10.1002/mas.10024. Mass Spectrom Rev. 2002. PMID: 12476441 Review.
-
Chemical cross-linking in the structural analysis of protein assemblies.Methods. 2018 Jul 15;144:53-63. doi: 10.1016/j.ymeth.2018.05.023. Epub 2018 May 30. Methods. 2018. PMID: 29857191 Free PMC article. Review.
-
Mapping protein interfaces by a trifunctional cross-linker combined with MALDI-TOF and ESI-FTICR mass spectrometry.J Am Soc Mass Spectrom. 2005 Dec;16(12):1921-31. doi: 10.1016/j.jasms.2005.07.020. Epub 2005 Oct 24. J Am Soc Mass Spectrom. 2005. PMID: 16246579
-
Chemical cross-linking and mass spectrometry for mapping three-dimensional structures of proteins and protein complexes.J Mass Spectrom. 2003 Dec;38(12):1225-37. doi: 10.1002/jms.559. J Mass Spectrom. 2003. PMID: 14696200
Cited by
-
RNA Chemical Labeling with Site-Specific, Relative Quantification by Mass Spectrometry for the Structural Study of a Neomycin-Sensing Riboswitch Aptamer Domain.Chempluschem. 2022 Nov;87(11):e202200256. doi: 10.1002/cplu.202200256. Epub 2022 Oct 11. Chempluschem. 2022. PMID: 36220343 Free PMC article.
-
Protein-RNA interactions: from mass spectrometry to drug discovery.Essays Biochem. 2023 Mar 29;67(2):175-186. doi: 10.1042/EBC20220177. Essays Biochem. 2023. PMID: 36866608 Free PMC article.
-
Chemical crosslinking extends and complements UV crosslinking in analysis of RNA/DNA nucleic acid-protein interaction sites by mass spectrometry.Nucleic Acids Res. 2025 Aug 11;53(15):gkaf727. doi: 10.1093/nar/gkaf727. Nucleic Acids Res. 2025. PMID: 40823814 Free PMC article.
-
B-CePs as cross-linking probes for the investigation of RNA higher-order structure.Nucleic Acids Res. 2021 Jul 9;49(12):6660-6672. doi: 10.1093/nar/gkab468. Nucleic Acids Res. 2021. PMID: 34125908 Free PMC article.
-
Investigating Protein-Nucleic Acid Binding Interactions with Diethylpyrocarbonate Covalent Labeling-Mass Spectrometry.J Am Soc Mass Spectrom. 2024 Oct 2;35(10):2272-2275. doi: 10.1021/jasms.4c00285. Epub 2024 Aug 29. J Am Soc Mass Spectrom. 2024. PMID: 39208253
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

