ITGA7 functions as a tumor suppressor and regulates migration and invasion in breast cancer
- PMID: 29760566
- PMCID: PMC5937492
- DOI: 10.2147/CMAR.S160379
ITGA7 functions as a tumor suppressor and regulates migration and invasion in breast cancer
Abstract
Background: Breast cancer is the most common malignancy in women and the underlying mechanism of breast cancer cell metastasis is still far from uncover. Integrin subunit alpha 7 (ITGA7) is a functioning protein. It has been detected in many malignancies. But the function of ITGA7 in breast cancer is not clear. Our aim is to explore ITGA7 expression and its role in breast cancer.
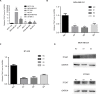
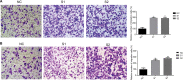
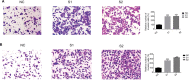
Methods: Real-time PCR was performed to determine ITGA7 expression in BC tissues and normal adjacent tissues. The specific functions of ITGA7 in breast cancer cell lines (MDA-MB-231 and BT-549) transfected with small interfering RNA were determined through migration, invasion assays. Western blot assays were performed to determine the expression of c-met and vimentin.
Results: ITGA7 was down-regulated in breast cancer tissues compared to the adjacent normal tissues (T:N =7.68±27.38: 41.01± 31.47, P<0.001) and this observation was consistent with the TCGA cohort (T:N =4.51±0.45:5.40±0.61, P<0.0001). In vitro experiments showed that knocking down ITGA7 significantly inhibited the migration and invasion of the breast cancer cell lines (MDA-MB-231 and BT-549). Meanwhile, knockdown of ITGA7 promoted c-met and vimentin expression, which may induce invasion and migration.
Conclusion: ITGA7 plays an important tumorigenic function and acts as a suppress gene in breast cancer. Our findings indicate that ITGA7 was the gene associated with breast cancer.
Keywords: ITGA7; breast cancer; invasion; migration.
Conflict of interest statement
Disclosure The authors report no conflicts of interest in this work.
Figures


References
-
- Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61(2):69–90. - PubMed
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. - PubMed
-
- Ahmed MI, Harvey JR, Kirby J, Ali S, Lennard TWJ. O-98 role of the chemokine receptor CXCR4 in breast cancer metastasis. Eur J Cancer Suppl. 2007;5(3):30.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

