The CRISPR tool kit for genome editing and beyond
- PMID: 29765029
- PMCID: PMC5953931
- DOI: 10.1038/s41467-018-04252-2
The CRISPR tool kit for genome editing and beyond
Abstract
CRISPR is becoming an indispensable tool in biological research. Once known as the bacterial immune system against invading viruses, the programmable capacity of the Cas9 enzyme is now revolutionizing diverse fields of medical research, biotechnology, and agriculture. CRISPR-Cas9 is no longer just a gene-editing tool; the application areas of catalytically impaired inactive Cas9, including gene regulation, epigenetic editing, chromatin engineering, and imaging, now exceed the gene-editing functionality of WT Cas9. Here, we will present a brief history of gene-editing tools and describe the wide range of CRISPR-based genome-targeting tools. We will conclude with future directions and the broader impact of CRISPR technologies.
Conflict of interest statement
The authors declare no competing interests.
Figures
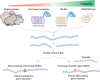



References
-
- Smith HO, Wilcox KW. A restriction enzyme from Hemophilus influenzae. I. Purification and general properties. J. Mol. Biol. 1970;51:379–391. - PubMed
-
- Kelly TJ, Jr., Smith HO. A restriction enzyme from Hemophilus influenzae. II. J. Mol. Biol. 1970;51:393–409. - PubMed
-
- Rothstein RJ. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. - PubMed
-
- Smithies O, Gregg RG, Boggs SS, Koralewski MA, Kucherlapati RS. Insertion of DNA sequences into the human chromosomal beta-globin locus by homologous recombination. Nature. 1985;317:230–234. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

