Comparative genomic analysis of multidrug-resistant Streptococcus pneumoniae isolates
- PMID: 29765237
- PMCID: PMC5939923
- DOI: 10.2147/IDR.S147858
Comparative genomic analysis of multidrug-resistant Streptococcus pneumoniae isolates
Abstract
Introduction: Multidrug resistance in Streptococcus pneumoniae has emerged as a serious problem to public health. A further understanding of the genetic diversity in antibiotic-resistant S. pneumoniae isolates is needed.
Methods: We conducted whole-genome resequencing for 25 pneumococcal strains isolated from children with different antimicrobial resistance profiles. Comparative analysis focus on detection of single-nucleotide polymorphisms (SNPs) and insertions and deletions (indels) was conducted. Moreover, phylogenetic analysis was applied to investigate the genetic relationship among these strains.
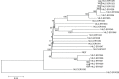
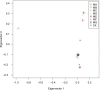
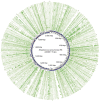
Results: The genome size of the isolates was ~2.1 Mbp, covering >90% of the total estimated size of the reference genome. The overall G+C% content was ~39.5%, and there were 2,200-2,400 open reading frames. All isolates with different drug resistance profiles harbored many indels (range 131-171) and SNPs (range 16,103-28,128). Genetic diversity analysis showed that the variation of different genes were associated with specific antibiotic resistance. Known antibiotic resistance genes (pbps, murMN, ciaH, rplD, sulA, and dpr) were identified, and new genes (regR, argH, trkH, and PTS-EII) closely related with antibiotic resistance were found, although these genes were primarily annotated with functions in virulence as well as carbohydrate and amino acid transport and metabolism. Phylogenetic analysis unambiguously indicated that isolates with different antibiotic resistance profiles harbored similar genetic backgrounds. One isolate, 14-LC.ER1025, showed a much weaker phylogenetic relationship with the other isolates, possibly caused by genomic variation.
Conclusion: In this study, although pneumococcal isolates had similar genetic backgrounds, strains were diverse at the genomic level. These strains exhibited distinct variations in their indel and SNP compositions associated with drug resistance.
Keywords: SNPs; Streptococcus pneumoniae; antimicrobial resistance; insertions/deletions; phylogenetic analysis; whole-genome sequencing.
Conflict of interest statement
Disclosure All authors report no potential conflicts of interest.
Figures

References
-
- World Health Organization Pneumococcal conjugate vaccine for childhood immunization: WHO position paper. Wkly Epidemiol Rec. 2007;82(12):93–104. - PubMed
-
- Kim SH, Song JH, Chung DR, et al. ANSORP Study Group Changing trends in antimicrobial resistance and serotypes of Streptococcus pneumoniae isolates in Asian countries: an Asian Network for Surveillance of Resistant Pathogens (ANSORP) study. Antimicrob Agents Chemother. 2012;56(3):1418–1426. - PMC - PubMed
-
- Cherazard R, Epstein M, Doan TL, Salim T, Bharti S, Smith MA. Antimicrobial resistant Streptococcus pneumoniae: prevalence, mechanisms, and clinical implications. Am J Ther. 2017;24(3):e361–e369. - PubMed
-
- Schweizer I, Blättner S, Maurer P, et al. New aspects of the interplay between penicillin binding proteins, murM, and the two-component system CiaRH of penicillin-resistant Streptococcus pneumoniae Serotype 19A isolates from Hungary. Antimicrob Agents Chemother. 2017;61(7):e00414–e00417. - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

