Crystal structures of Lymphocytic choriomeningitis virus endonuclease domain complexed with diketo-acid ligands
- PMID: 29765612
- PMCID: PMC5947727
- DOI: 10.1107/S2052252518001021
Crystal structures of Lymphocytic choriomeningitis virus endonuclease domain complexed with diketo-acid ligands
Abstract
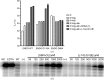
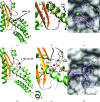
The Arenaviridae family, together with the Bunyaviridae and Orthomyxoviridae families, is one of the three negative-stranded RNA viral families that encode an endonuclease in their genome. The endonuclease domain is at the N-terminus of the L protein, a multifunctional protein that includes the RNA-dependent RNA polymerase. The synthesis of mRNA in arenaviruses is a process that is primed by capped nucleotides that are 'stolen' from the cellular mRNA by the endonuclease domain in cooperation with other domains of the L protein. This molecular mechanism has been demonstrated previously for the endonuclease of the prototype Lymphocytic choriomeningitis virus (LCMV). However, the mode of action of this enzyme is not fully understood as the original structure did not contain catalytic metal ions. The pivotal role played by the cap-snatching process in the life cycle of the virus and the highly conserved nature of the endonuclease domain make it a target of choice for the development of novel antiviral therapies. Here, the binding affinities of two diketo-acid (DKA) compounds (DPBA and L-742,001) for the endonuclease domain of LCMV were evaluated using biophysical methods. X-ray structures of the LCMV endonuclease domain with catalytic ions in complex with these two compounds were determined, and their efficacies were assessed in an in vitro endonuclease-activity assay. Based on these data and computational simulation, two new DKAs were synthesized. The LCMV endonuclease domain exhibits a good affinity for these DKAs, making them a good starting point for the design of arenavirus endonuclease inhibitors. In addition to providing the first example of an X-ray structure of an arenavirus endonuclease incorporating a ligand, this study provides a proof of concept that the design of optimized inhibitors against the arenavirus endonuclease is possible.
Keywords: Arenaviridae; LCMV; Lymphocytic choriomeningitis virus; compound optimization; diketo acids; endonucleases; metal chelation.
Figures



References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

