The Subgingival Microbiome of Periodontal Pockets With Different Probing Depths in Chronic and Aggressive Periodontitis: A Pilot Study
- PMID: 29765908
- PMCID: PMC5938363
- DOI: 10.3389/fcimb.2018.00124
The Subgingival Microbiome of Periodontal Pockets With Different Probing Depths in Chronic and Aggressive Periodontitis: A Pilot Study
Abstract
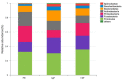
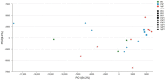
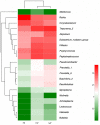
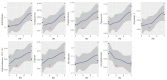
Periodontitis is a kind of infectious disease initiated by colonization of subgingival periodontal pathogens, which cause destruction of tooth-supporting tissues, and is a predominant threat to oral health as the most common cause of loss of teeth. The aim of this pilot study was to characterize the subgingival bacterial biodiversity of periodontal pockets with different probing depths in patients with different forms of periodontitis. Twenty-one subgingival plaque samples were collected from three patients with chronic periodontitis (ChP), three patients with aggressive periodontitis (AgP) and three periodontally healthy subjects (PH). Each patient with periodontitis was sampled at three sites, at different probing depths (PDs, one each at 4 mm, 5-6 mm, and ≥ 7 mm). Using 16S rRNA gene high-throughput sequencing and bioinformatic analysis, we found that subgingival communities in health and periodontitis samples largely differed. Meanwhile, Acholeplasma, Fretibacterium, Porphyromonas, Peptococcus, Treponema_2, Defluviitaleaceae_UCG_011, Filifactor, and Mycoplasma increased with the deepening of the pockets in ChP, whilst only Corynebacterium was negatively associated with PD. In AgP, Corynebacterium and Klebsiella were positively associated with PD, while Serratia, Pseudoramibacter, Defluviitaleaceae_UCG_011, and Desulfobulbus were negatively associated with PD. And among these two groups, Corynebacterium shifted differently. Moreover, in subgingival plaque, the unweighted UniFrac distances between samples from pockets with different PD in the same patients were significantly lower than those from pockets within the same PD category from different patients. This study demonstrated the shift of the subgingival microbiome in individual teeth sites during disease development. Within the limitation of the relative small sample size, this pilot study shed new light on the dynamic relationship between the extent of periodontal destruction and the subgingival microbiome.
Keywords: 16S rDNA; aggressive periodontitis; chronic periodontitis; high-throughput nucleotide sequencing; subgingival microbiome.
Figures




Similar articles
-
Comparison of Subgingival and Buccal Mucosa Microbiome in Chronic and Aggressive Periodontitis: A Pilot Study.Front Cell Infect Microbiol. 2019 Mar 11;9:53. doi: 10.3389/fcimb.2019.00053. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 30915280 Free PMC article.
-
Peri-implant and periodontal microbiome diversity in aggressive periodontitis patients: a pilot study.Clin Oral Implants Res. 2017 May;28(5):558-570. doi: 10.1111/clr.12834. Epub 2016 May 11. Clin Oral Implants Res. 2017. PMID: 27170047
-
[Comparison of subgingival microbial profile of aggressive periodontitis, chronic periodontitis and periodontally healthy individuals].Zhonghua Kou Qiang Yi Xue Za Zhi. 2020 Jul 9;55(7):466-474. doi: 10.3760/cma.j.cn112144-20200413-00207. Zhonghua Kou Qiang Yi Xue Za Zhi. 2020. PMID: 32634885 Chinese.
-
Influence of periodontal surgery on the subgingival microbiome-A systematic review and meta-analysis.J Periodontal Res. 2023 Apr;58(2):308-324. doi: 10.1111/jre.13092. Epub 2023 Jan 4. J Periodontal Res. 2023. PMID: 36597817
-
Aggressive Periodontitis: microbes and host response, who to blame?Virulence. 2015;6(3):223-8. doi: 10.4161/21505594.2014.986407. Virulence. 2015. PMID: 25654663 Free PMC article. Review.
Cited by
-
Prevalence of herpes virus in chronic periodontitis patients with and without type 2 diabetes mellitus: A clinico-microbiological study.J Oral Maxillofac Pathol. 2021 Jan-Apr;25(1):141-146. doi: 10.4103/jomfp.JOMFP_154_20. Epub 2021 May 14. J Oral Maxillofac Pathol. 2021. PMID: 34349425 Free PMC article.
-
The Subgingival Microbial Composition in Health and Periodontitis with Different Probing Depths.Microorganisms. 2025 Apr 17;13(4):930. doi: 10.3390/microorganisms13040930. Microorganisms. 2025. PMID: 40284767 Free PMC article.
-
Oral microbiota analyses of paediatric Saudi population reveals signatures of dental caries.BMC Oral Health. 2023 Nov 27;23(1):935. doi: 10.1186/s12903-023-03448-3. BMC Oral Health. 2023. PMID: 38012587 Free PMC article.
-
Development and characterization of an oral microbiome transplant among Australians for the treatment of dental caries and periodontal disease: A study protocol.PLoS One. 2021 Nov 29;16(11):e0260433. doi: 10.1371/journal.pone.0260433. eCollection 2021. PLoS One. 2021. PMID: 34843568 Free PMC article.
-
Periodontal pockets: A potential reservoir for SARS-CoV-2?Med Hypotheses. 2020 Oct;143:109907. doi: 10.1016/j.mehy.2020.109907. Epub 2020 May 30. Med Hypotheses. 2020. PMID: 32504927 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

