Long reads: their purpose and place
- PMID: 29767702
- PMCID: PMC6061690
- DOI: 10.1093/hmg/ddy177
Long reads: their purpose and place
Abstract
In recent years long-read technologies have moved from being a niche and specialist field to a point of relative maturity likely to feature frequently in the genomic landscape. Analogous to next generation sequencing, the cost of sequencing using long-read technologies has materially dropped whilst the instrument throughput continues to increase. Together these changes present the prospect of sequencing large numbers of individuals with the aim of fully characterizing genomes at high resolution. In this article, we will endeavour to present an introduction to long-read technologies showing: what long reads are; how they are distinct from short reads; why long reads are useful and how they are being used. We will highlight the recent developments in this field, and the applications and potential of these technologies in medical research, and clinical diagnostics and therapeutics.
Figures
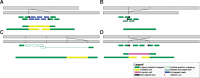
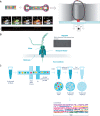

References
-
- Sanger F., Coulson A. (1975) A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. J. Mol. Biol., 94, 441–448. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

