Identification of a Recurrent LMO7-BRAF Fusion in Papillary Thyroid Carcinoma
- PMID: 29768105
- PMCID: PMC5994666
- DOI: 10.1089/thy.2017.0258
Identification of a Recurrent LMO7-BRAF Fusion in Papillary Thyroid Carcinoma
Abstract
Background: The BRAFV600E mutation is the most common driver in papillary thyroid carcinoma (PTC) tumors. In recent years, gene fusions have also been recognized as important drivers of cancer in PTC. Previous studies have suggested that thyroid tumors with fusion genes frequently display an aggressive course. These observations prompted further exploration of gene fusions in PTC tumors. The aim was to search for previously unrecognized gene fusions using thyroid tissue samples from PTC patients.
Methods: Gene fusions were analyzed in RNA sequencing data obtained from 12 PTC tumors and paired unaffected thyroid tissue samples. Candidate fusions were further filtered and validated using reverse transcriptase polymerase chain reaction, Sanger sequencing, and fluorescence in situ hybridization. An Ohio cohort of 148 PTC tumor samples was screened for a LMO7-BRAF fusion and the BRAFV600E mutation. Functional assays were performed to assess the LMO7-BRAF fusion.
Results: Two coding fusions (CCDC6-RET and LMO7-BRAF) were found in one tumor sample each. The novel LMO7-BRAF fusion was validated by reverse transcriptase polymerase chain reaction and fluorescence in situ hybridization. The LMO7-BRAF fusion was a recurrent somatic alteration with a frequency of 2.0% (3/148) in PTC tumors, while the BRAFV600E point mutation was found in 63.5% (94/148) of tumors. Enforced expression of LMO7-BRAF fusion protein stimulated endogenous ERK1/2 phosphorylation and promoted anchorage independent cell growth to an extent similar to BRAFV600E.
Conclusions: A novel fusion gene, LMO7-BRAF, was identified in PTC tumors. The results indicate that the LMO7-BRAF fusion behaves as an oncogenic alteration. This observation expands the spectrum of fusion genes involving kinases in thyroid cancer.
Keywords: BRAFV600E; LMO7–BRAF; fusion gene; papillary thyroid carcinoma.
Conflict of interest statement
The authors have no conflicts of interest to declare.
Figures

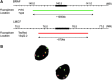

Similar articles
-
AGK-BRAF gene fusion is a recurrent event in sporadic pediatric thyroid carcinoma.Cancer Med. 2016 Jul;5(7):1535-41. doi: 10.1002/cam4.698. Epub 2016 Mar 31. Cancer Med. 2016. PMID: 27037835 Free PMC article.
-
RET, NTRK, ALK, BRAF, and MET Fusions in a Large Cohort of Pediatric Papillary Thyroid Carcinomas.Thyroid. 2020 Dec;30(12):1771-1780. doi: 10.1089/thy.2019.0802. Epub 2020 Jul 1. Thyroid. 2020. PMID: 32495721
-
Molecular Profiling of Papillary Thyroid Carcinoma in Korea with a High Prevalence of BRAFV600E Mutation.Thyroid. 2017 Jun;27(6):802-810. doi: 10.1089/thy.2016.0547. Epub 2017 Apr 21. Thyroid. 2017. PMID: 28293988
-
[BRAF initiating mutations in the papillary thyroid carcinoma].Endokrynol Pol. 2006 Jul-Aug;57(4):438-44. Endokrynol Pol. 2006. PMID: 17006850 Review. Polish.
-
Evidence that one subset of anaplastic thyroid carcinomas are derived from papillary carcinomas due to BRAF and p53 mutations.Cancer. 2005 Jun 1;103(11):2261-8. doi: 10.1002/cncr.21073. Cancer. 2005. PMID: 15880523 Review.
Cited by
-
Integrated analysis of colorectal cancer metastasis identifies characteristics of tumor cell during metastasis.Gastroenterol Rep (Oxf). 2024 May 30;12:goae055. doi: 10.1093/gastro/goae055. eCollection 2024. Gastroenterol Rep (Oxf). 2024. PMID: 38818308 Free PMC article.
-
Lmo7 recruits myosin II heavy chain to regulate actomyosin contractility and apical domain size in Xenopus ectoderm.Development. 2022 May 15;149(10):dev200236. doi: 10.1242/dev.200236. Epub 2022 May 16. Development. 2022. PMID: 35451459 Free PMC article.
-
Radiofrequency ablation is an effective treatment for Bethesda III thyroid nodules without genetic alterations.Eur Thyroid J. 2024 May 13;13(3):e240020. doi: 10.1530/ETJ-24-0020. Print 2024 Jun 1. Eur Thyroid J. 2024. PMID: 38657647 Free PMC article.
-
LMO7 as an Unrecognized Factor Promoting Pancreatic Cancer Progression and Metastasis.Front Cell Dev Biol. 2021 Mar 8;9:647387. doi: 10.3389/fcell.2021.647387. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 33763427 Free PMC article.
-
Novel Gene Signatures Predictive of Patient Recurrence-Free Survival and Castration Resistance in Prostate Cancer.Cancers (Basel). 2021 Feb 22;13(4):917. doi: 10.3390/cancers13040917. Cancers (Basel). 2021. PMID: 33671634 Free PMC article.
References
-
- Davies L, Welch HG. 2014. Current thyroid cancer trends in the United States. JAMA Otolaryngol Head Neck Surg 140:317–322 - PubMed
-
- La Vecchia C, Negri E. 2017. Thyroid cancer: the thyroid cancer epidemic—overdiagnosis or a real increase? Nat Rev Endocrinol 13:318–319 - PubMed
-
- Siegel RL, Miller KD, Jemal A. 2017. Cancer statistics, 2017. CA Cancer J Clin 67:7–30 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

