ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase
- PMID: 29769718
- PMCID: PMC5973781
- DOI: 10.1038/s41586-018-0137-8
ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase
Erratum in
-
Publisher Correction: ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase.Nature. 2018 Aug;560(7720):E35. doi: 10.1038/s41586-018-0271-3. Nature. 2018. PMID: 29925958
Abstract
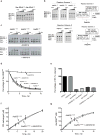
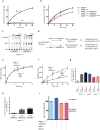
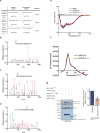
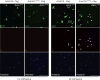
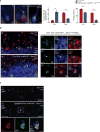
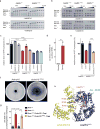
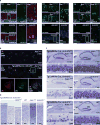
Editing domains of aminoacyl tRNA synthetases correct tRNA charging errors to maintain translational fidelity. A mutation in the editing domain of alanyl tRNA synthetase (AlaRS) in Aars sti mutant mice results in an increase in the production of serine-mischarged tRNAAla and the degeneration of cerebellar Purkinje cells. Here, using positional cloning, we identified Ankrd16, a gene that acts epistatically with the Aars sti mutation to attenuate neurodegeneration. ANKRD16, a vertebrate-specific protein that contains ankyrin repeats, binds directly to the catalytic domain of AlaRS. Serine that is misactivated by AlaRS is captured by the lysine side chains of ANKRD16, which prevents the charging of serine adenylates to tRNAAla and precludes serine misincorporation in nascent peptides. The deletion of Ankrd16 in the brains of Aarssti/sti mice causes widespread protein aggregation and neuron loss. These results identify an amino-acid-accepting co-regulator of tRNA synthetase editing as a new layer of the machinery that is essential to the prevention of severe pathologies that arise from defects in editing.
Figures








Similar articles
-
Substrate specificity and catalysis by the editing active site of Alanyl-tRNA synthetase from Escherichia coli.Biochemistry. 2011 Mar 8;50(9):1474-82. doi: 10.1021/bi1013535. Epub 2011 Jan 31. Biochemistry. 2011. PMID: 21241052 Free PMC article.
-
Editing-defective tRNA synthetase causes protein misfolding and neurodegeneration.Nature. 2006 Sep 7;443(7107):50-5. doi: 10.1038/nature05096. Epub 2006 Aug 13. Nature. 2006. PMID: 16906134
-
Paradox of mistranslation of serine for alanine caused by AlaRS recognition dilemma.Nature. 2009 Dec 10;462(7274):808-12. doi: 10.1038/nature08612. Nature. 2009. PMID: 20010690 Free PMC article.
-
The uniqueness of AlaRS and its human disease connections.RNA Biol. 2021 Nov;18(11):1501-1511. doi: 10.1080/15476286.2020.1861803. Epub 2020 Dec 23. RNA Biol. 2021. PMID: 33317386 Free PMC article. Review.
-
Trans-editing by aminoacyl-tRNA synthetase-like editing domains.Enzymes. 2020;48:69-115. doi: 10.1016/bs.enz.2020.07.002. Epub 2020 Sep 8. Enzymes. 2020. PMID: 33837712 Review.
Cited by
-
Tryptophanylation of insulin receptor by WARS attenuates insulin signaling.Cell Mol Life Sci. 2024 Jan 12;81(1):25. doi: 10.1007/s00018-023-05082-2. Cell Mol Life Sci. 2024. PMID: 38212570 Free PMC article.
-
The mechanism of β-N-methylamino-l-alanine inhibition of tRNA aminoacylation and its impact on misincorporation.J Biol Chem. 2020 Jan 31;295(5):1402-1410. doi: 10.1074/jbc.RA119.011714. Epub 2019 Dec 20. J Biol Chem. 2020. PMID: 31862734 Free PMC article.
-
Leucyl-tRNA synthetase is a tumour suppressor in breast cancer and regulates codon-dependent translation dynamics.Nat Cell Biol. 2022 Mar;24(3):307-315. doi: 10.1038/s41556-022-00856-5. Epub 2022 Mar 14. Nat Cell Biol. 2022. PMID: 35288656 Free PMC article.
-
Overlapping spectrums: The clinicogenetic commonalities between Charcot-Marie-Tooth and other neurodegenerative diseases.Brain Res. 2020 Jan 15;1727:146532. doi: 10.1016/j.brainres.2019.146532. Epub 2019 Oct 31. Brain Res. 2020. PMID: 31678418 Free PMC article. Review.
-
Defects in aminoacyl-tRNA synthetase cause partial B and T cell immunodeficiency.Cell Mol Life Sci. 2022 Jan 23;79(2):87. doi: 10.1007/s00018-021-04122-z. Cell Mol Life Sci. 2022. PMID: 35067747 Free PMC article.
References
-
- Ling J, Reynolds N, Ibba M. Aminoacyl-tRNA synthesis and translational quality control. Annu. Rev. Microbiol. 2009;63:61–78. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

