TP53 gain-of-function mutation promotes inflammation in glioblastoma
- PMID: 29786075
- PMCID: PMC6370770
- DOI: 10.1038/s41418-018-0126-3
TP53 gain-of-function mutation promotes inflammation in glioblastoma
Abstract
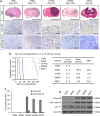
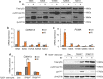
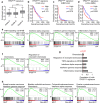
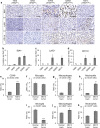
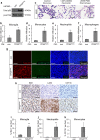
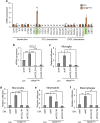
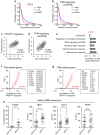
Glioblastoma (GBM), the most severe and common brain tumor in adults, is characterized by multiple somatic mutations and aberrant activation of inflammatory responses. Immune cell infiltration and subsequent inflammation cause tumor growth and resistance to therapy. Somatic loss-of-function mutations in the gene encoding tumor suppressor protein p53 (TP53) are frequently observed in various cancers. However, numerous studies suggest that TP53 regulates malignant phenotypes by gain-of-function (GOF) mutations. Here we demonstrate that a TP53 GOF mutation promotes inflammation in GBM. Ectopic expression of a TP53 GOF mutant induced transcriptomic changes, which resulted in enrichment of gene signatures related to inflammation and chemotaxis. Bioinformatics analyses revealed that a gene signature, upregulated by the TP53 GOF mutation, is associated with progression and shorter overall survival in GBM. We also observed significant correlations between the TP53 GOF mutation signature and inflammation in the clinical database of GBM and other cancers. The TP53 GOF mutant showed upregulated C-C motif chemokine ligand 2 (CCL2) and tumor necrosis factor alpha (TNFA) expression via nuclear factor kappa B (NFκB) signaling, consequently increasing microglia and monocyte-derived immune cell infiltration. Additionally, TP53 GOF mutation and CCL2 and TNFA expression correlated positively with tumor-associated immunity in patients with GBM. Taken together, our findings suggest that the TP53 GOF mutation plays a crucial role in inflammatory responses, thereby deteriorating prognostic outcomes in patients with GBM.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
Similar articles
-
Biallelic TP53 gain of function mutations in rapidly progressing solid tumors.Cancer Genet. 2018 Apr;222-223:20-24. doi: 10.1016/j.cancergen.2018.02.001. Epub 2018 Feb 24. Cancer Genet. 2018. PMID: 29666004
-
Elevation of the TP53 isoform Δ133p53β in glioblastomas: an alternative to mutant p53 in promoting tumor development.J Pathol. 2018 Sep;246(1):77-88. doi: 10.1002/path.5111. Epub 2018 Jul 31. J Pathol. 2018. PMID: 29888503 Free PMC article.
-
The role of TP53 gain-of-function mutation in multifocal glioblastoma.J Neurooncol. 2020 Mar;147(1):37-47. doi: 10.1007/s11060-019-03318-5. Epub 2020 Jan 30. J Neurooncol. 2020. PMID: 32002804 Free PMC article.
-
Integrative analysis of aberrant Wnt signaling in hepatitis B virus-related hepatocellular carcinoma.World J Gastroenterol. 2015 May 28;21(20):6317-28. doi: 10.3748/wjg.v21.i20.6317. World J Gastroenterol. 2015. PMID: 26034368 Free PMC article. Review.
-
p53 and disease: when the guardian angel fails.Cell Death Differ. 2006 Jun;13(6):1017-26. doi: 10.1038/sj.cdd.4401913. Cell Death Differ. 2006. PMID: 16557268 Review.
Cited by
-
Targeting drug resistance in glioblastoma (Review).Int J Oncol. 2024 Aug;65(2):80. doi: 10.3892/ijo.2024.5668. Epub 2024 Jul 12. Int J Oncol. 2024. PMID: 38994761 Free PMC article. Review.
-
Multiomics integration reveals the effect of Orexin A on glioblastoma.Front Pharmacol. 2023 Jan 20;14:1096159. doi: 10.3389/fphar.2023.1096159. eCollection 2023. Front Pharmacol. 2023. PMID: 36744263 Free PMC article.
-
Network Pharmacology of Yougui Pill Combined with Buzhong Yiqi Decoction for the Treatment of Sexual Dysfunction.Evid Based Complement Alternat Med. 2019 Nov 11;2019:1243743. doi: 10.1155/2019/1243743. eCollection 2019. Evid Based Complement Alternat Med. 2019. PMID: 31814838 Free PMC article.
-
Bioinformatics analysis of DNMT1 expression and its role in head and neck squamous cell carcinoma prognosis.Sci Rep. 2021 Jan 26;11(1):2267. doi: 10.1038/s41598-021-81971-5. Sci Rep. 2021. PMID: 33500531 Free PMC article.
-
Multiple augmented reduced rank regression for pan-cancer analysis.Biometrics. 2024 Jan 29;80(1):ujad002. doi: 10.1093/biomtc/ujad002. Biometrics. 2024. PMID: 38281771 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

