Uncoupling of sgRNAs from their associated barcodes during PCR amplification of combinatorial CRISPR screens
- PMID: 29799876
- PMCID: PMC5969736
- DOI: 10.1371/journal.pone.0197547
Uncoupling of sgRNAs from their associated barcodes during PCR amplification of combinatorial CRISPR screens
Abstract
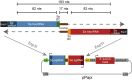
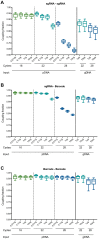
Many implementations of pooled screens in mammalian cells rely on linking an element of interest to a barcode, with the latter subsequently quantitated by next generation sequencing. However, substantial uncoupling between these paired elements during lentiviral production has been reported, especially as the distance between elements increases. We detail that PCR amplification is another major source of uncoupling, and becomes more pronounced with increased amounts of DNA template molecules and PCR cycles. To lessen uncoupling in systems that use paired elements for detection, we recommend minimizing the distance between elements, using low and equal template DNA inputs for plasmid and genomic DNA during PCR, and minimizing the number of PCR cycles. We also present a vector design for conducting combinatorial CRISPR screens that enables accurate barcode-based detection with a single short sequencing read and minimal uncoupling.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Maricque BB, Dougherty JD, Cohen BA. A genome-integrated massively parallel reporter assay reveals DNA sequence determinants of cis-regulatory activity in neural cells. Nucleic Acids Res. 2017;45: e16 doi: 10.1093/nar/gkw942 - DOI - PMC - PubMed
-
- Patwardhan RP, Lee C, Litvin O, Young DL, Pe’er D, Shendure J. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Nat Biotechnol. 2009;27: 1173–1175. doi: 10.1038/nbt.1589 - DOI - PMC - PubMed
-
- Melnikov A, Zhang X, Rogov P, Wang L, Mikkelsen TS. Massively parallel reporter assays in cultured mammalian cells. J Vis Exp. 2014; doi: 10.3791/51719 - DOI - PMC - PubMed
-
- Doench JG. Am I ready for CRISPR? A user’s guide to genetic screens. Nat Rev Genet. 2018;19: 67–80. doi: 10.1038/nrg.2017.97 - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

