Genomic analyses of unique carbohydrate and phytohormone metabolism in the macroalga Gracilariopsis lemaneiformis (Rhodophyta)
- PMID: 29801464
- PMCID: PMC5970526
- DOI: 10.1186/s12870-018-1309-2
Genomic analyses of unique carbohydrate and phytohormone metabolism in the macroalga Gracilariopsis lemaneiformis (Rhodophyta)
Abstract
Background: Red algae are economically valuable for food and in industry. However, their genomic information is limited, and the genomic data of only a few species of red algae have been sequenced and deposited recently. In this study, we annotated a draft genome of the macroalga Gracilariopsis lemaneiformis (Gracilariales, Rhodophyta).
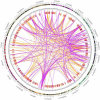
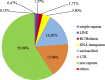
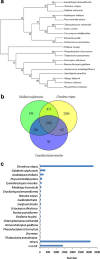
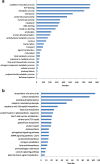
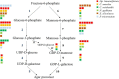
Results: The entire 88.98 Mb genome of Gp. lemaneiformis 981 was generated from 13,825 scaffolds (≥500 bp) with an N50 length of 30,590 bp, accounting for approximately 91% of this algal genome. A total of 38.73 Mb of scaffold sequences were repetitive, and 9281 protein-coding genes were predicted. A phylogenomic analysis of 20 genomes revealed the relationship among the Chromalveolata, Rhodophyta, Chlorophyta and higher plants. Homology analysis indicated phylogenetic proximity between Gp. lemaneiformis and Chondrus crispus. The number of enzymes related to the metabolism of carbohydrates, including agar, glycoside hydrolases, glycosyltransferases, was abundant. In addition, signaling pathways associated with phytohormones such as auxin, salicylic acid and jasmonates are reported for the first time for this alga.
Conclusion: We sequenced and analyzed a draft genome of the red alga Gp. lemaneiformis, and revealed its carbohydrate metabolism and phytohormone signaling characteristics. This work will be helpful in research on the functional and comparative genomics of the order Gracilariales and will enrich the genomic information on marine algae.
Keywords: Carbohydrate metabolism; Genomic analysis; Gracilariopsis lemaneiformis; Phytohormone signaling.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Complete mitochondrial genome of agar-producing red alga Gracilariopsis chorda (Gracilariales).Mitochondrial DNA. 2014 Oct;25(5):339-41. doi: 10.3109/19401736.2013.800502. Epub 2013 Jun 21. Mitochondrial DNA. 2014. PMID: 23789772
-
Comparative transcriptional profiling of Gracilariopsis lemaneiformis in response to salicylic acid- and methyl jasmonate-mediated heat resistance.PLoS One. 2017 May 2;12(5):e0176531. doi: 10.1371/journal.pone.0176531. eCollection 2017. PLoS One. 2017. PMID: 28464018 Free PMC article.
-
Genome survey sequencing and genetic background characterization of Gracilariopsis lemaneiformis (Rhodophyta) based on next-generation sequencing.PLoS One. 2013 Jul 16;8(7):e69909. doi: 10.1371/journal.pone.0069909. Print 2013. PLoS One. 2013. PMID: 23875008 Free PMC article.
-
JAZ repressors and the orchestration of phytohormone crosstalk.Trends Plant Sci. 2012 Jan;17(1):22-31. doi: 10.1016/j.tplants.2011.10.006. Epub 2011 Nov 21. Trends Plant Sci. 2012. PMID: 22112386 Review.
-
Evolution of jasmonate biosynthesis and signaling mechanisms.J Exp Bot. 2017 Mar 1;68(6):1323-1331. doi: 10.1093/jxb/erw470. J Exp Bot. 2017. PMID: 28007954 Review.
Cited by
-
Identification of Indicator Genes for Agar Accumulation in Gracilariopsis lemaneiformis (Rhodophyta).Int J Mol Sci. 2024 Apr 23;25(9):4606. doi: 10.3390/ijms25094606. Int J Mol Sci. 2024. PMID: 38731824 Free PMC article.
-
Physiological, Transcriptome, and Metabolome Analyses Reveal the Tolerance to Cu Toxicity in Red Macroalgae Gracilariopsis lemaneiformis.Int J Mol Sci. 2024 Apr 27;25(9):4770. doi: 10.3390/ijms25094770. Int J Mol Sci. 2024. PMID: 38731988 Free PMC article.
-
Heterozygous Single Nucleotide Polymorphic Loci in Haploid Gametophytes of Gracilariopsis lemaneiformis (Rhodophyta).Front Genet. 2019 Dec 6;10:1256. doi: 10.3389/fgene.2019.01256. eCollection 2019. Front Genet. 2019. PMID: 31921299 Free PMC article.
-
The Genome of the Marine Alga Ulva compressa (Chlorophyta) Reveals Protein-Coding Genes with Similarity to Plants and Green Microalgae, but Also to Animal, Bacterial, and Fungal Genes.Int J Mol Sci. 2022 Jun 30;23(13):7279. doi: 10.3390/ijms23137279. Int J Mol Sci. 2022. PMID: 35806287 Free PMC article.
-
A survey of the full-length transcriptome of Gracilariopsis lemaneiformis using single-molecule long-read sequencing.BMC Plant Biol. 2022 Dec 19;22(1):597. doi: 10.1186/s12870-022-03992-0. BMC Plant Biol. 2022. PMID: 36536287 Free PMC article.
References
-
- Bird CJ, Ragan MA, Critchley AT, Rice EL, Gutell RR. Molecular relationships among the Gracilariaceae (Rhodophyta): further observations on some undetermined species. Eur J Phycol. 1994;29:195–202. doi: 10.1080/09670269400650641. - DOI
-
- Gurgel CFD, Liao LM, Fredericq S, Hommersand MH. Systematics of Gracilariopsis Dawson (Gracilariales, Rhodophyta) based on rbcL sequence analysis and morphological evidence. J Phycol. 2003;39:154–171. doi: 10.1046/j.1529-8817.2003.02046.x. - DOI
-
- Yu J, Yang YF. Physiological and biochemical response of seaweed Gracilaria lemaneiformis to concentration changes of N and P. J Exp Mar Biol Ecol. 2008;367:142–148. doi: 10.1016/j.jembe.2008.09.009. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

