Atg9 establishes Atg2-dependent contact sites between the endoplasmic reticulum and phagophores
- PMID: 29848619
- PMCID: PMC6080931
- DOI: 10.1083/jcb.201710116
Atg9 establishes Atg2-dependent contact sites between the endoplasmic reticulum and phagophores
Abstract
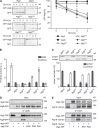
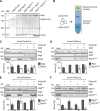
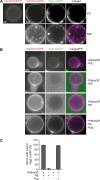
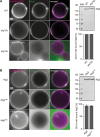
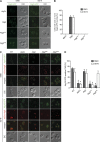
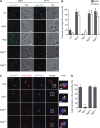
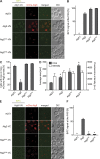
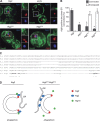
The autophagy-related (Atg) proteins play a key role in the formation of autophagosomes, the hallmark of autophagy. The function of the cluster composed by Atg2, Atg18, and transmembrane Atg9 is completely unknown despite their importance in autophagy. In this study, we provide insights into the molecular role of these proteins by identifying and characterizing Atg2 point mutants impaired in Atg9 binding. We show that Atg2 associates to autophagosomal membranes through lipid binding and independently from Atg9. Its interaction with Atg9, however, is key for Atg2 confinement to the growing phagophore extremities and subsequent association of Atg18. Assembly of the Atg9-Atg2-Atg18 complex is important to establish phagophore-endoplasmic reticulum (ER) contact sites. In turn, disruption of the Atg2-Atg9 interaction leads to an aberrant topological distribution of both Atg2 and ER contact sites on forming phagophores, which severely impairs autophagy. Altogether, our data shed light in the interrelationship between Atg9, Atg2, and Atg18 and highlight the possible functional relevance of the phagophore-ER contact sites in phagophore expansion.
© 2018 Gómez-Sánchez et al.
Figures


References
-
- Axe E.L., Walker S.A., Manifava M., Chandra P., Roderick H.L., Habermann A., Griffiths G., and Ktistakis N.T.. 2008. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 182:685–701. 10.1083/jcb.200803137 - DOI - PMC - PubMed
-
- Bakula D., Müller A.J., Zuleger T., Takacs Z., Franz-Wachtel M., Thost A.K., Brigger D., Tschan M.P., Frickey T., Robenek H., et al. . 2017. WIPI3 and WIPI4 β-propellers are scaffolds for LKB1-AMPK-TSC signalling circuits in the control of autophagy. Nat. Commun. 8:15637 10.1038/ncomms15637 - DOI - PMC - PubMed
-
- Brachmann C.B., Davies A., Cost G.J., Caputo E., Li J., Hieter P., and Boeke J.D.. 1998. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast. 14:115–132. 10.1002/(SICI)1097-0061(19980130)14:2<115::AID-YEA204>3.0.CO;2-2 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

