Analysis of Genetic Variation Indicates DNA Shape Involvement in Purifying Selection
- PMID: 29850830
- PMCID: PMC6063282
- DOI: 10.1093/molbev/msy099
Analysis of Genetic Variation Indicates DNA Shape Involvement in Purifying Selection
Abstract
Noncoding DNA sequences, which play various roles in gene expression and regulation, are under evolutionary pressure. Gene regulation requires specific protein-DNA binding events, and our previous studies showed that both DNA sequence and shape readout are employed by transcription factors (TFs) to achieve DNA binding specificity. By investigating the shape-disrupting properties of single nucleotide polymorphisms (SNPs) in human regulatory regions, we established a link between disruptive local DNA shape changes and loss of specific TF binding. Furthermore, we described cases where disease-associated SNPs may alter TF binding through DNA shape changes. This link led us to hypothesize that local DNA shape within and around TF binding sites is under selection pressure. To verify this hypothesis, we analyzed SNP data derived from 216 natural strains of Drosophila melanogaster. Comparing SNPs located in functional and nonfunctional regions within experimentally validated cis-regulatory modules (CRMs) from D. melanogaster that are active in the blastoderm stage of development, we found that SNPs within functional regions tended to cause smaller DNA shape variations. Furthermore, SNPs with higher minor allele frequency were more likely to result in smaller DNA shape variations. The same analysis based on a large number of SNPs in putative CRMs of the D. melanogaster genome derived from DNase I accessibility data confirmed these observations. Taken together, our results indicate that common SNPs in functional regions tend to maintain DNA shape, whereas shape-disrupting SNPs are more likely to be eliminated through purifying selection.
Figures

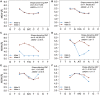
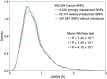
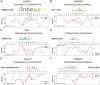
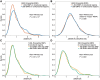
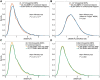
Similar articles
-
Cis-regulatory variations: a study of SNPs around genes showing cis-linkage in segregating mouse populations.BMC Genomics. 2006 Sep 15;7:235. doi: 10.1186/1471-2164-7-235. BMC Genomics. 2006. PMID: 16978413 Free PMC article.
-
Analysis of genomic variation in non-coding elements using population-scale sequencing data from the 1000 Genomes Project.Nucleic Acids Res. 2011 Sep 1;39(16):7058-76. doi: 10.1093/nar/gkr342. Epub 2011 May 19. Nucleic Acids Res. 2011. PMID: 21596777 Free PMC article.
-
Translating natural genetic variation to gene expression in a computational model of the Drosophila gap gene regulatory network.PLoS One. 2017 Sep 12;12(9):e0184657. doi: 10.1371/journal.pone.0184657. eCollection 2017. PLoS One. 2017. PMID: 28898266 Free PMC article.
-
Transcription factor family-specific DNA shape readout revealed by quantitative specificity models.Mol Syst Biol. 2017 Feb 6;13(2):910. doi: 10.15252/msb.20167238. Mol Syst Biol. 2017. PMID: 28167566 Free PMC article.
-
Single nucleotide polymorphism in transcriptional regulatory regions and expression of environmentally responsive genes.Toxicol Appl Pharmacol. 2005 Sep 1;207(2 Suppl):84-90. doi: 10.1016/j.taap.2004.09.024. Toxicol Appl Pharmacol. 2005. PMID: 16002116 Review.
Cited by
-
RiceSNP-BST: a deep learning framework for predicting biotic stress-associated SNPs in rice.Brief Bioinform. 2024 Sep 23;25(6):bbae599. doi: 10.1093/bib/bbae599. Brief Bioinform. 2024. PMID: 39562160 Free PMC article.
-
Landscape of DNA binding signatures of myocyte enhancer factor-2B reveals a unique interplay of base and shape readout.Nucleic Acids Res. 2020 Sep 4;48(15):8529-8544. doi: 10.1093/nar/gkaa642. Nucleic Acids Res. 2020. PMID: 32738045 Free PMC article.
-
TFBSshape: an expanded motif database for DNA shape features of transcription factor binding sites.Nucleic Acids Res. 2020 Jan 8;48(D1):D246-D255. doi: 10.1093/nar/gkz970. Nucleic Acids Res. 2020. PMID: 31665425 Free PMC article.
-
Genetic Variants in Transcription Factor Binding Sites in Humans: Triggered by Natural Selection and Triggers of Diseases.Int J Mol Sci. 2021 Apr 18;22(8):4187. doi: 10.3390/ijms22084187. Int J Mol Sci. 2021. PMID: 33919522 Free PMC article. Review.
-
REDfly: An Integrated Knowledgebase for Insect Regulatory Genomics.Insects. 2022 Jul 11;13(7):618. doi: 10.3390/insects13070618. Insects. 2022. PMID: 35886794 Free PMC article. Review.
References
-
- Abelson JF, Kwan KY, O’Roak BJ, Baek DY, Stillman AA, Morgan TM, Mathews CA, Pauls DL, Rasin MR, Gunel M, et al. 2005. Sequence variants in SLITRK1 are associated with Tourette’s syndrome. Science 3105746: 317–320. - PubMed
-
- Alipanahi B, Delong A, Weirauch MT, Frey BJ.. 2015. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat Biotechnol. 338: 831–838. - PubMed
-
- Andolfatto P. 2005. Adaptive evolution of non-coding DNA in Drosophila. Nature 4377062: 1149–1152. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

