Evolutionary convergence in lignin-degrading enzymes
- PMID: 29866821
- PMCID: PMC6016776
- DOI: 10.1073/pnas.1802555115
Evolutionary convergence in lignin-degrading enzymes
Abstract
The resurrection of ancestral enzymes of now-extinct organisms (paleogenetics) is a developing field that allows the study of evolutionary hypotheses otherwise impossible to be tested. In the present study, we target fungal peroxidases that play a key role in lignin degradation, an essential process in the carbon cycle and often a limiting step in biobased industries. Ligninolytic peroxidases are secreted by wood-rotting fungi, the origin of which was recently established in the Carboniferous period associated with the appearance of these enzymes. These first peroxidases were not able to degrade lignin directly and used diffusible metal cations to attack its phenolic moiety. The phylogenetic analysis of the peroxidases of Polyporales, the order in which most extant wood-rotting fungi are included, suggests that later in evolution these enzymes would have acquired the ability to degrade nonphenolic lignin using a tryptophanyl radical interacting with the bulky polymer at the surface of the enzyme. Here, we track this powerful strategy for lignin degradation as a phenotypic trait in fungi and show that it is not an isolated event in the evolution of Polyporales. Using ancestral enzyme resurrection, we study the molecular changes that led to the appearance of the same surface oxidation site in two distant peroxidase lineages. By characterization of the resurrected enzymes, we demonstrate convergent evolution at the amino acid level during the evolution of these fungi and track the different changes leading to phylogenetically distant ligninolytic peroxidases from ancestors lacking the ability to degrade nonphenolic lignin.
Keywords: Polyporales; ancestral enzyme resurrection; convergent evolution; fungal peroxidases; lignin.
Copyright © 2018 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


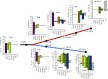


References
-
- Martínez AT, Ruiz-Dueñas FJ, Martínez MJ, Del Río JC, Gutiérrez A. Enzymatic delignification of plant cell wall: From nature to mill. Curr Opin Biotechnol. 2009;20:348–357. - PubMed
-
- Hammel KE, Cullen D. Role of fungal peroxidases in biological ligninolysis. Curr Opin Plant Biol. 2008;11:349–355. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

