Genome-wide characterization and expression analysis of GRAS gene family in pepper (Capsicum annuum L.)
- PMID: 29868257
- PMCID: PMC5983004
- DOI: 10.7717/peerj.4796
Genome-wide characterization and expression analysis of GRAS gene family in pepper (Capsicum annuum L.)
Abstract
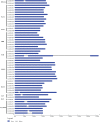
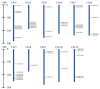
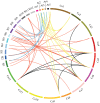
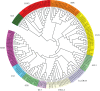
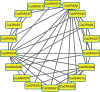
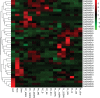
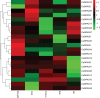
Plant-specific GRAS transcription factors regulate various biological processes in plant growth, development and stress responses. However, this important gene family was not fully characterized in pepper (Capsicum annuum L.), an economically important vegetable crop. Here, a total of 50 CaGRAS members were identified in pepper genome and renamed by their respective chromosomal distribution. Genomic organization revealed that most CaGRAS genes (84%) have no intron. Phylogenetic analysis divided pepper CaGRAS members into 10 subfamilies, with each having distinct conserved domains and functions. For the expansion of the GRAS genes in pepper, segmental duplication contributed more than tandem duplication did. Gene expression analysis in various tissues demonstrated that most of CaGRAS genes exhibited a tissue- and development stage-specific expression pattern, uncovering their potential functions in pepper growth and development. Moreover, 21 CaGRAS genes were differentially expressed under cold, drought, salt and gibberellin acid (GA) treatments, indicating that they may implicated in plant response to abiotic stress. Notably, GA responsive cis-elements were detected in the promoter regions of the majority of CaGRAS genes, suggesting that CaGRAS may involve in signal cross-talking. The first comprehensive analysis of GRAS gene family in pepper genome by this study provide insights into understanding the GRAS-mediated regulation network, benefiting the genetic improvements in pepper and some other relative plants.
Keywords: Abiotic stress; Duplication; GRAS genes; Gene expression; Pepper.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Genome-wide identification and expression analysis of the GRAS gene family in response to drought stress in chickpea (Cicer arietinum L.).3 Biotech. 2022 Mar;12(3):64. doi: 10.1007/s13205-021-03104-z. Epub 2022 Feb 9. 3 Biotech. 2022. PMID: 35186661 Free PMC article.
-
Genome wide identification and expression analysis of pepper C2H2 zinc finger transcription factors in response to anthracnose pathogen Colletotrichum truncatum.3 Biotech. 2021 Mar;11(3):118. doi: 10.1007/s13205-020-02601-x. Epub 2021 Feb 8. 3 Biotech. 2021. PMID: 33747699 Free PMC article.
-
Genome-Wide Analyses of the NAC Transcription Factor Gene Family in Pepper (Capsicum annuum L.): Chromosome Location, Phylogeny, Structure, Expression Patterns, Cis-Elements in the Promoter, and Interaction Network.Int J Mol Sci. 2018 Mar 29;19(4):1028. doi: 10.3390/ijms19041028. Int J Mol Sci. 2018. PMID: 29596349 Free PMC article.
-
The SnRK2 family in pepper (Capsicum annuum L.): genome-wide identification and expression analyses during fruit development and under abiotic stress.Genes Genomics. 2020 Oct;42(10):1117-1130. doi: 10.1007/s13258-020-00968-y. Epub 2020 Jul 31. Genes Genomics. 2020. PMID: 32737808 Review.
-
Final report on the safety assessment of capsicum annuum extract, capsicum annuum fruit extract, capsicum annuum resin, capsicum annuum fruit powder, capsicum frutescens fruit, capsicum frutescens fruit extract, capsicum frutescens resin, and capsaicin.Int J Toxicol. 2007;26 Suppl 1:3-106. doi: 10.1080/10915810601163939. Int J Toxicol. 2007. PMID: 17365137 Review.
Cited by
-
Characterization of the GRAS gene family reveals their contribution to the high adaptability of wheat.PeerJ. 2021 Feb 23;9:e10811. doi: 10.7717/peerj.10811. eCollection 2021. PeerJ. 2021. PMID: 33665016 Free PMC article.
-
Genome-wide identification, expression analysis and functional study of the GRAS gene family in Tartary buckwheat (Fagopyrum tataricum).BMC Plant Biol. 2019 Aug 6;19(1):342. doi: 10.1186/s12870-019-1951-3. BMC Plant Biol. 2019. PMID: 31387526 Free PMC article.
-
Genome-wide identification of GRAS genes in Brachypodium distachyon and functional characterization of BdSLR1 and BdSLRL1.BMC Genomics. 2019 Aug 6;20(1):635. doi: 10.1186/s12864-019-5985-6. BMC Genomics. 2019. PMID: 31387534 Free PMC article.
-
Genome-Wide Identification, Expression and Stress Analysis of the GRAS Gene Family in Phoebe bournei.Plants (Basel). 2023 May 21;12(10):2048. doi: 10.3390/plants12102048. Plants (Basel). 2023. PMID: 37653964 Free PMC article.
-
Genome-Wide Identification and Analysis of the GRAS Transcription Factor Gene Family in Theobroma cacao.Genes (Basel). 2022 Dec 24;14(1):57. doi: 10.3390/genes14010057. Genes (Basel). 2022. PMID: 36672798 Free PMC article.
References
-
- Abarca D, Pizarro A, Hernandez I, Sanchez C, Solana SP, Del Amo A, Carneros E, Diaz-Sala C. The GRAS gene family in pine: transcript expression patterns associated with the maturation-related decline of competence to form adventitious roots. BMC Plant Biology. 2014;14(1):1–19. doi: 10.1186/s12870-014-0354-8. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

