Targeting CAND1 promotes caspase-8/RIP1-dependent apoptosis in liver cancer cells
- PMID: 29887951
- PMCID: PMC5992550
Targeting CAND1 promotes caspase-8/RIP1-dependent apoptosis in liver cancer cells
Abstract
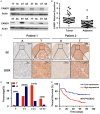
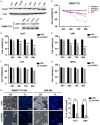
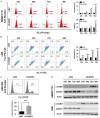
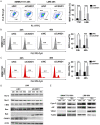
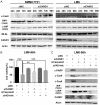
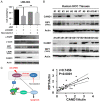
Cullin-associated NEDD8-dissociated 1 (CAND1) plays a vital role in regulating the activity of Cullin-RING ubiquitin ligases (CRLs), which are frequently dysregulated in cancer. However, the role of CAND1 in hepatocellular carcinoma (HCC) remains unknown. Here, we found that CAND1 was overexpressed in HCC tissues compared to corresponding adjacent liver tissues (71.7% vs 16.7%); high expression of CAND1 was associated with poor overall survival (40.7 vs 57.3 months, P=0.0013); and CAND1 was an independent risk factor for the prognosis of HCC patients (N=138, P=0.018). Functional studies revealed that CAND1 knockdown efficiently suppressed the proliferation of liver cancer cells by activating caspase-8-dependent mitochondrial apoptosis. We also observed a mutual activation loop between caspase-8 and Receptor Interacting Protein 1 (RIP1), which amplified CAND1 knockdown-induced apoptotic signals in the cells. Furthermore, RIP1 inhibitor Necrostatin-1 eliminated the activation of caspase-8. In conclusion, our study pioneered in reporting high CAND1 expression as a predictor of poor prognosis for HCC patients. CAND1 silencing suppressed HCC cell proliferation by inducing caspase-8/RIP1-dependent apoptosis. These findings supported that CAND1 could be a new therapeutic target for liver cancer.
Keywords: Cullin-associated NEDD8-dissociated 1 (CAND1); Receptor Interacting Protein 1 (RIP1); apoptosis; caspase-8; hepatocellular carcinoma (HCC).
Conflict of interest statement
None.
Figures
Similar articles
-
Cullin-associated and neddylation-dissociated 1 regulate reprogramming of lipid metabolism through SKP1-Cullin-1-F-boxFBXO11 -mediated heterogeneous nuclear ribonucleoprotein A2/B1 ubiquitination and promote hepatocellular carcinoma.Clin Transl Med. 2023 Oct;13(10):e1443. doi: 10.1002/ctm2.1443. Clin Transl Med. 2023. PMID: 37837399 Free PMC article.
-
CAND1-dependent control of cullin 1-RING Ub ligases is essential for adipogenesis.Biochim Biophys Acta. 2013 May;1833(5):1078-84. doi: 10.1016/j.bbamcr.2013.01.005. Epub 2013 Jan 14. Biochim Biophys Acta. 2013. PMID: 23328082
-
Drosophila Cand1 regulates Cullin3-dependent E3 ligases by affecting the neddylation of Cullin3 and by controlling the stability of Cullin3 and adaptor protein.Dev Biol. 2010 Oct 15;346(2):247-57. doi: 10.1016/j.ydbio.2010.07.031. Epub 2010 Aug 4. Dev Biol. 2010. PMID: 20691177
-
Assembly and Regulation of CRL Ubiquitin Ligases.Adv Exp Med Biol. 2020;1217:33-46. doi: 10.1007/978-981-15-1025-0_3. Adv Exp Med Biol. 2020. PMID: 31898220 Review.
-
Regulation of cullin RING ligases.Annu Rev Plant Biol. 2008;59:467-89. doi: 10.1146/annurev.arplant.58.032806.104011. Annu Rev Plant Biol. 2008. PMID: 18444905 Review.
Cited by
-
The Impact of Cand1 in Prostate Cancer.Cancers (Basel). 2020 Feb 12;12(2):428. doi: 10.3390/cancers12020428. Cancers (Basel). 2020. PMID: 32059441 Free PMC article.
-
Bioinformatics Analysis of the Prognostic Significance of CAND1 in ERα-Positive Breast Cancer.Diagnostics (Basel). 2022 Sep 27;12(10):2327. doi: 10.3390/diagnostics12102327. Diagnostics (Basel). 2022. PMID: 36292029 Free PMC article.
-
Integration of Fungus-Specific CandA-C1 into a Trimeric CandA Complex Allowed Splitting of the Gene for the Conserved Receptor Exchange Factor of CullinA E3 Ubiquitin Ligases in Aspergilli.mBio. 2019 Jun 18;10(3):e01094-19. doi: 10.1128/mBio.01094-19. mBio. 2019. PMID: 31213557 Free PMC article.
-
Ubiquitin Proteasome Pathway Transcriptome in Epithelial Ovarian Cancer.Cancers (Basel). 2021 May 28;13(11):2659. doi: 10.3390/cancers13112659. Cancers (Basel). 2021. PMID: 34071321 Free PMC article. Review.
-
Targeting caspase-8: a new strategy for combating hepatocellular carcinoma.Front Immunol. 2024 Dec 12;15:1501659. doi: 10.3389/fimmu.2024.1501659. eCollection 2024. Front Immunol. 2024. PMID: 39726605 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
