The Odd "RB" Phage-Identification of Arabinosylation as a New Epigenetic Modification of DNA in T4-Like Phage RB69
- PMID: 29890699
- PMCID: PMC6024577
- DOI: 10.3390/v10060313
The Odd "RB" Phage-Identification of Arabinosylation as a New Epigenetic Modification of DNA in T4-Like Phage RB69
Abstract
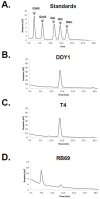
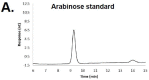
In bacteriophages related to T4, hydroxymethylcytosine (hmC) is incorporated into the genomic DNA during DNA replication and is then further modified to glucosyl-hmC by phage-encoded glucosyltransferases. Previous studies have shown that RB69 shares a core set of genes with T4 and relatives. However, unlike the other “RB” phages, RB69 is unable to recombine its DNA with T4 or with the other “RB” isolates. In addition, despite having homologs to the T4 enzymes used to synthesize hmC, RB69 has no identified homolog to known glucosyltransferase genes. In this study we sought to understand the basis for RB69’s behavior using high-pH anion exchange chromatography (HPAEC) and mass spectrometry. Our analyses identified a novel phage epigenetic DNA sugar modification in RB69 DNA, which we have designated arabinosyl-hmC (ara-hmC). We sought a putative glucosyltranserase responsible for this novel modification and determined that RB69 also has a novel transferase gene, ORF003c, that is likely responsible for the arabinosyl-specific modification. We propose that ara-hmC was responsible for RB69 being unable to participate in genetic exchange with other hmC-containing T-even phages, and for its described incipient speciation. The RB69 ara-hmC also likely protects its DNA from some anti-phage type-IV restriction endonucleases. Several T4-related phages, such as E. coli phage JS09 and Shigella phage Shf125875 have homologs to RB69 ORF003c, suggesting the ara-hmC modification may be relatively common in T4-related phages, highlighting the importance of further work to understand the role of this modification and the biochemical pathway responsible for its production.
Keywords: RB69; T4 phage; anion exchange chromatography; arabinosyl-hmC (ara-hmC); epigenetic modification; glucosyl-hmC (ghmC); glucosyltransferase transferase; hydroxymethylcytosine (hmC); matrix assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry; restriction.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- JKutter E., Stidham T., Guttman B., Kutter E., Batts D., Peterson S., Djavakhishvili T., Arisaka F., Mesyanzhinov V., Ruger W., et al. Genomic map of bacteriophage T4. In: Karam J.D., editor. Molecular Biology of Bacteriophage T4. ASM Press; Washington, DC, USA: 1994. pp. 491–519.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

