Rectal and Naris Swabs: Practical and Informative Samples for Analyzing the Microbiota of Critically Ill Patients
- PMID: 29898981
- PMCID: PMC6001609
- DOI: 10.1128/mSphere.00219-18
Rectal and Naris Swabs: Practical and Informative Samples for Analyzing the Microbiota of Critically Ill Patients
Erratum in
-
Erratum for Bansal et al., "Rectal and Naris Swabs: Practical and Informative Samples for Analyzing the Microbiota of Critically Ill Patients".mSphere. 2018 Jun 27;3(3):e00328-18. doi: 10.1128/mSphere.00328-18. eCollection 2018 May-Jun. mSphere. 2018. PMID: 31329809 Free PMC article.
Abstract
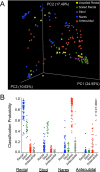
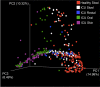
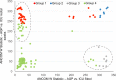
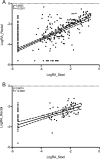
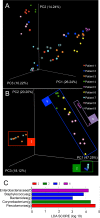
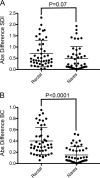
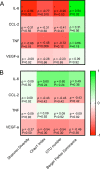
Commensal microbiota are immunomodulatory, and their pathological perturbation can affect the risk and outcomes of infectious and inflammatory diseases. Consequently, the human microbiota is an emerging diagnostic and therapeutic target in critical illness. In this study, we compared four sample types-rectal, naris, and antecubital swabs and stool samples-for 16S rRNA gene microbiota sequencing in intensive care unit (ICU) patients. Stool samples were obtained in only 31% of daily attempts, while swabs were reliably obtained (≥97% of attempts). Swabs were compositionally distinct by anatomical site, and rectal swabs identified within-patient temporal trends in microbiota composition. Rectal swabs from ICU patients demonstrated differences from healthy stool similar to those observed in comparing stool samples from ICU patients to those from the same healthy controls. Rectal swabs are a useful complement to other sample types for analysis of the intestinal microbiota in critical illness, particularly when obtaining stool may not be feasible or practical.IMPORTANCE Perturbation of the microbiome has been correlated with various infectious and inflammatory diseases and is common in critically ill patients. Stool is typically used to sample the microbiota in human observational studies; however, it is often unavailable for collection from critically ill patients, reducing its utility as a sample type to study this population. Our research identified alternatives to stool for sampling the microbiota during critical illness. Rectal and naris swabs were practical alternatives for use in these patients, as they were observed to be more reliably obtained than stool, were suitable for culture-independent analysis, and successfully captured within- and between-patient microbiota differences.
Keywords: 16S RNA; critical illness; human microbiome.
Copyright © 2018 Bansal et al.
Figures
References
-
- Vincent J-L, Rello J, Marshall J, Silva E, Anzueto A, Martin CD, Moreno R, Lipman J, Gomersall C, Sakr Y, Reinhart K; EPIC II Group of Investigators . 2009. International study of the prevalence and outcomes of infection in intensive care units. JAMA 302:2323. doi: 10.1001/jama.2009.1754. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
