Systematic Identification, Characterization, and Conservation of Adjacent-Gene Coregulation in the Budding Yeast Saccharomyces cerevisiae
- PMID: 29898982
- PMCID: PMC6001612
- DOI: 10.1128/mSphere.00220-18
Systematic Identification, Characterization, and Conservation of Adjacent-Gene Coregulation in the Budding Yeast Saccharomyces cerevisiae
Abstract
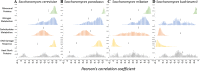
It is essential that cells orchestrate gene expression for the specific niche that they occupy, and this often requires coordination of the expression of large sets of genes. There are multiple regulatory systems that exist for modulation of gene expression, including the adjacent-gene coregulation of the rRNA and ribosome biogenesis and ribosomal protein families. Both gene families exhibit a nonrandom genomic distribution, often clustered directly adjacent to another member of the same family, which results in a tighter transcriptional coordination among adjacent paired genes than that of the unpaired genes within each regulon and can result in a shared promoter that coordinates expression of the pairs. This nonrandom genomic distribution has been seen in a few functionally related gene families, and many of these functional pairings are conserved across divergent fungal lineages. To date, the significance of these observations has not been extended in a systematic way to characterize how prevalent the role of adjacent-gene coregulation is in transcriptional regulation. In the present study, we systematically analyzed the transcriptional coherence of the functional pairs compared to the singletons within all gene families defined by the Gene Ontology Slim designation, using Saccharomyces cerevisiae as a model system, finding that clusters exhibit a tighter transcriptional correlation under specific contexts. We found that the longer a functional pairing is conserved the tighter its response to broad stress and nutritional responses, that roughly 25% of gene families exhibit a nonrandom genomic distribution, and that many of these clusters are conserved. This suggests that adjacent-gene coregulation is a widespread, yet underappreciated, transcriptional mechanism.IMPORTANCE The spatial positioning of genes throughout the genome arrangement can alter their expression in many eukaryotic organisms. Often this results in a genomic context-specific effect on transcription. One example of this is through the clustering of functionally related genes, which results in adjacent-gene coregulation in the budding yeast Saccharomyces cerevisiae In the present study, we set out to systematically characterize the prevalence of this phenomenon, finding the genomic organization of functionally related genes into clusters is a characteristic of myriad gene families. These arrangements are found in many evolutionarily divergent fungi and thus represent a widespread, yet underappreciated, layer of transcriptional regulation.
Keywords: Saccharomyces cerevisiae; adjacent-gene coregulation; gene expression; spatial positioning.
Copyright © 2018 Eldabagh et al.
Figures

Similar articles
-
Functionally Related Genes Cluster into Genomic Regions That Coordinate Transcription at a Distance in Saccharomyces cerevisiae.mSphere. 2019 Mar 13;4(2):e00063-19. doi: 10.1128/mSphere.00063-19. mSphere. 2019. PMID: 30867326 Free PMC article.
-
Dissecting the cis and trans elements that regulate adjacent-gene coregulation in Saccharomyces cerevisiae.Eukaryot Cell. 2014 Jun;13(6):738-48. doi: 10.1128/EC.00317-13. Epub 2014 Apr 4. Eukaryot Cell. 2014. PMID: 24706020 Free PMC article.
-
The adjacent positioning of co-regulated gene pairs is widely conserved across eukaryotes.BMC Genomics. 2012 Oct 10;13:546. doi: 10.1186/1471-2164-13-546. BMC Genomics. 2012. PMID: 23051624 Free PMC article.
-
Coexpression, coregulation, and cofunctionality of neighboring genes in eukaryotic genomes.Genomics. 2008 Mar;91(3):243-8. doi: 10.1016/j.ygeno.2007.11.002. Epub 2007 Dec 21. Genomics. 2008. PMID: 18082363 Review.
-
Yeast transcriptional regulatory mechanisms.Annu Rev Genet. 1995;29:651-74. doi: 10.1146/annurev.ge.29.120195.003251. Annu Rev Genet. 1995. PMID: 8825489 Review.
Cited by
-
Systematic Analysis of Functionally Related Gene Clusters in the Opportunistic Pathogen, Candida albicans.Microorganisms. 2021 Jan 28;9(2):276. doi: 10.3390/microorganisms9020276. Microorganisms. 2021. PMID: 33525750 Free PMC article.
-
Genomic clustering within functionally related gene families in Ascomycota fungi.Comput Struct Biotechnol J. 2020 Oct 29;18:3267-3277. doi: 10.1016/j.csbj.2020.10.020. eCollection 2020. Comput Struct Biotechnol J. 2020. PMID: 33209211 Free PMC article. Review.
-
Trans-acting genetic variation affects the expression of adjacent genes.Genetics. 2021 Mar 31;217(3):iyaa051. doi: 10.1093/genetics/iyaa051. Genetics. 2021. PMID: 33789351 Free PMC article.
-
Functionally Related Genes Cluster into Genomic Regions That Coordinate Transcription at a Distance in Saccharomyces cerevisiae.mSphere. 2019 Mar 13;4(2):e00063-19. doi: 10.1128/mSphere.00063-19. mSphere. 2019. PMID: 30867326 Free PMC article.
-
Transcriptional control of ribosome biogenesis in yeast: links to growth and stress signals.Biochem Soc Trans. 2021 Aug 27;49(4):1589-1599. doi: 10.1042/BST20201136. Biochem Soc Trans. 2021. PMID: 34240738 Free PMC article. Review.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
