Intracellular biosynthesis of lipids and cholesterol by Scap and Insig in mesenchymal cells regulates long bone growth and chondrocyte homeostasis
- PMID: 29899135
- PMCID: PMC6053657
- DOI: 10.1242/dev.162396
Intracellular biosynthesis of lipids and cholesterol by Scap and Insig in mesenchymal cells regulates long bone growth and chondrocyte homeostasis
Abstract
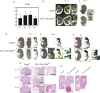
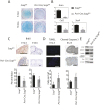
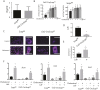
During enchondral ossification, mesenchymal cells express genes regulating the intracellular biosynthesis of cholesterol and lipids. Here, we have investigated conditional deletion of Scap or of Insig1 and Insig2 (Scap inhibits intracellular biosynthesis and Insig proteins activate intracellular biosynthesis). Mesenchymal condensation and chondrogenesis was disrupted in mice lacking Scap in mesenchymal progenitors, whereas mice lacking the Insig genes in mesenchymal progenitors had short limbs, but normal chondrogenesis. Mice lacking Scap in chondrocytes showed severe dwarfism, with ectopic hypertrophic cells, whereas deletion of Insig genes in chondrocytes caused a mild dwarfism and shortening of the hypertrophic zone. In vitro studies showed that intracellular cholesterol in chondrocytes can derive from exogenous and endogenous sources, but that exogenous sources cannot completely overcome the phenotypic effect of Scap deficiency. Genes encoding cholesterol biosynthetic proteins are regulated by Hedgehog (Hh) signaling, and Hh signaling is also regulated by intracellular cholesterol in chondrocytes, suggesting a feedback loop in chondrocyte differentiation. Precise regulation of intracellular biosynthesis is required for chondrocyte homeostasis and long bone growth, and these data support pharmacological modulation of cholesterol biosynthesis as a therapy for select cartilage pathologies.
Keywords: Cholesterol; Chondrocyte; Enchodral; Insig; Scap.
© 2018. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures




Similar articles
-
Regulation of Cholesterol Homeostasis by Hedgehog Signaling in Osteoarthritic Cartilage.Arthritis Rheumatol. 2016 Jan;68(1):127-37. doi: 10.1002/art.39337. Arthritis Rheumatol. 2016. PMID: 26315393 Free PMC article.
-
The hedgehog target Vlk genetically interacts with Gli3 to regulate chondrocyte differentiation during mouse long bone development.Differentiation. 2013 Apr-Jun;85(4-5):121-30. doi: 10.1016/j.diff.2013.03.002. Epub 2013 Jun 19. Differentiation. 2013. PMID: 23792766
-
Sterol-regulated ubiquitination and degradation of Insig-1 creates a convergent mechanism for feedback control of cholesterol synthesis and uptake.Cell Metab. 2006 Jan;3(1):15-24. doi: 10.1016/j.cmet.2005.11.014. Cell Metab. 2006. PMID: 16399501
-
Regulation of bone development and extracellular matrix protein genes by RUNX2.Cell Tissue Res. 2010 Jan;339(1):189-95. doi: 10.1007/s00441-009-0832-8. Epub 2009 Aug 1. Cell Tissue Res. 2010. PMID: 19649655 Review.
-
Interaction of growth factors regulating chondrocyte differentiation in the developing embryo.Osteoarthritis Cartilage. 2001;9 Suppl A:S109-17. Osteoarthritis Cartilage. 2001. PMID: 11680674 Review.
Cited by
-
Acute hyperlipidemia has transient effects on large-scale bone regeneration in male mice.Sci Rep. 2024 Oct 27;14(1):25610. doi: 10.1038/s41598-024-76992-9. Sci Rep. 2024. PMID: 39463386 Free PMC article.
-
Intracellular cholesterol biosynthesis in enchondroma and chondrosarcoma.JCI Insight. 2019 Apr 30;5(11):e127232. doi: 10.1172/jci.insight.127232. JCI Insight. 2019. PMID: 31039139 Free PMC article.
-
Role of Metabolism in Bone Development and Homeostasis.Int J Mol Sci. 2020 Nov 26;21(23):8992. doi: 10.3390/ijms21238992. Int J Mol Sci. 2020. PMID: 33256181 Free PMC article. Review.
-
Enchondromatosis and Growth Plate Development.Curr Osteoporos Rep. 2021 Feb;19(1):40-49. doi: 10.1007/s11914-020-00639-7. Epub 2020 Dec 11. Curr Osteoporos Rep. 2021. PMID: 33306166 Free PMC article. Review.
-
Metabolic Pathways and Targets in Chondrosarcoma.Front Oncol. 2021 Dec 6;11:772263. doi: 10.3389/fonc.2021.772263. eCollection 2021. Front Oncol. 2021. PMID: 34938658 Free PMC article. Review.
References
-
- Ayciriex S., Regazzetti A., Gaudin M., Prost E., Dargère D., Massicot F., Auzeil N. and Laprévote O. (2012). Development of a novel method for quantification of sterols and oxysterols by UPLC-ESI-HRMS: application to a neuroinflammation rat model. Anal. Bioanal. Chem. 404, 3049-3059. 10.1007/s00216-012-6396-6 - DOI - PubMed
-
- Bornig H. and Geyer G. (1974). Staining of cholesterol with the fluorescent antibiotic “filipin”. Acta Histochem. 50, 110-115. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

