Modulating Enzyme Activity by Altering Protein Dynamics with Solvent
- PMID: 29901984
- PMCID: PMC6172137
- DOI: 10.1021/acs.biochem.8b00424
Modulating Enzyme Activity by Altering Protein Dynamics with Solvent
Abstract
Optimal enzyme activity depends on a number of factors, including structure and dynamics. The role of enzyme structure is well recognized; however, the linkage between protein dynamics and enzyme activity has given rise to a contentious debate. We have developed an approach that uses an aqueous mixture of organic solvent to control the functionally relevant enzyme dynamics (without changing the structure), which in turn modulates the enzyme activity. Using this approach, we predicted that the hydride transfer reaction catalyzed by the enzyme dihydrofolate reductase (DHFR) from Escherichia coli in aqueous mixtures of isopropanol (IPA) with water will decrease by ∼3 fold at 20% (v/v) IPA concentration. Stopped-flow kinetic measurements find that the pH-independent khydride rate decreases by 2.2 fold. X-ray crystallographic enzyme structures show no noticeable differences, while computational studies indicate that the transition state and electrostatic effects were identical for water and mixed solvent conditions; quasi-elastic neutron scattering studies show that the dynamical enzyme motions are suppressed. Our approach provides a unique avenue to modulating enzyme activity through changes in enzyme dynamics. Further it provides vital insights that show the altered motions of DHFR cause significant changes in the enzyme's ability to access its functionally relevant conformational substates, explaining the decreased khydride rate. This approach has important implications for obtaining fundamental insights into the role of rate-limiting dynamics in catalysis and as well as for enzyme engineering.
Figures
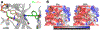


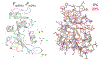

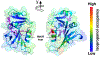
References
-
- Benkovic SJ, and Hammes-Schiffer S (2006) Biochemistry - Enzyme motions inside and out, Science 312, 208–209. - PubMed
-
- Boehr DD, Dyson HJ, and Wright PE (2006) An NMR perspective on enzyme dynamics, Chemical Reviews 106, 3055–3079. - PubMed
-
- Henzler-Wildman K, and Kern D (2007) Dynamic personalities of proteins, Nature 450, 964–972. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

