The Spectrum of Replication Errors in the Absence of Error Correction Assayed Across the Whole Genome of Escherichia coli
- PMID: 29907648
- PMCID: PMC6063229
- DOI: 10.1534/genetics.117.300515
The Spectrum of Replication Errors in the Absence of Error Correction Assayed Across the Whole Genome of Escherichia coli
Abstract
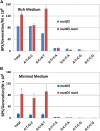
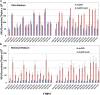
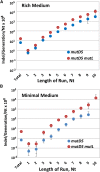
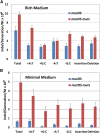
When the DNA polymerase that replicates the Escherichia coli chromosome, DNA polymerase III, makes an error, there are two primary defenses against mutation: proofreading by the ϵ subunit of the holoenzyme and mismatch repair. In proofreading-deficient strains, mismatch repair is partially saturated and the cell's response to DNA damage, the SOS response, may be partially induced. To investigate the nature of replication errors, we used mutation accumulation experiments and whole-genome sequencing to determine mutation rates and mutational spectra across the entire chromosome of strains deficient in proofreading, mismatch repair, and the SOS response. We report that a proofreading-deficient strain has a mutation rate 4000-fold greater than wild-type strains. While the SOS response may be induced in these cells, it does not contribute to the mutational load. Inactivating mismatch repair in a proofreading-deficient strain increases the mutation rate another 1.5-fold. DNA polymerase has a bias for converting G:C to A:T base pairs, but proofreading reduces the impact of these mutations, helping to maintain the genomic G:C content. These findings give an unprecedented view of how polymerase and error-correction pathways work together to maintain E. coli's low mutation rate of 1 per 1000 generations.
Keywords: DNA proofreading; DNA replication fidelity; mismatch repair; mutation accumulation; mutation hotspots.
Copyright © 2018 by the Genetics Society of America.
Figures
Similar articles
-
Genetic requirements and mutational specificity of the Escherichia coli SOS mutator activity.J Bacteriol. 1997 Dec;179(23):7435-45. doi: 10.1128/jb.179.23.7435-7445.1997. J Bacteriol. 1997. PMID: 9393709 Free PMC article.
-
Escherichia coli mutator (Delta)polA is defective in base mismatch correction: the nature of in vivo DNA replication errors.J Mol Biol. 2005 Aug 12;351(2):299-308. doi: 10.1016/j.jmb.2005.06.014. J Mol Biol. 2005. PMID: 16005896
-
Determinants of Base-Pair Substitution Patterns Revealed by Whole-Genome Sequencing of DNA Mismatch Repair Defective Escherichia coli.Genetics. 2018 Aug;209(4):1029-1042. doi: 10.1534/genetics.118.301237. Epub 2018 Jun 15. Genetics. 2018. PMID: 29907647 Free PMC article.
-
DNA replication fidelity in Escherichia coli: a multi-DNA polymerase affair.FEMS Microbiol Rev. 2012 Nov;36(6):1105-21. doi: 10.1111/j.1574-6976.2012.00338.x. Epub 2012 Apr 5. FEMS Microbiol Rev. 2012. PMID: 22404288 Free PMC article. Review.
-
Translesion synthesis in Escherichia coli: lessons from the NarI mutation hot spot.DNA Repair (Amst). 2007 Jul 1;6(7):1032-41. doi: 10.1016/j.dnarep.2007.02.021. Epub 2007 Apr 2. DNA Repair (Amst). 2007. PMID: 17403618 Review.
Cited by
-
Experimental evolution under different nutritional conditions changes the genomic architecture and virulence of Acinetobacter baumannii.Commun Biol. 2024 Oct 5;7(1):1274. doi: 10.1038/s42003-024-06978-w. Commun Biol. 2024. PMID: 39369115 Free PMC article.
-
Effect of mismatch repair on the mutational footprint of the bacterial SOS mutator activity.DNA Repair (Amst). 2021 Jul;103:103130. doi: 10.1016/j.dnarep.2021.103130. Epub 2021 May 9. DNA Repair (Amst). 2021. PMID: 33991871 Free PMC article.
-
Mutational Signatures in Wild Type Escherichia coli Strains Reveal Predominance of DNA Polymerase Errors.Genome Biol Evol. 2024 Apr 2;16(4):evae035. doi: 10.1093/gbe/evae035. Genome Biol Evol. 2024. PMID: 38401265 Free PMC article.
-
In vivo evolution of an emerging zoonotic bacterial pathogen in an immunocompromised human host.Nat Commun. 2021 Jul 23;12(1):4495. doi: 10.1038/s41467-021-24668-7. Nat Commun. 2021. PMID: 34301946 Free PMC article.
-
Antibiotic perseverance increases the risk of resistance development.Proc Natl Acad Sci U S A. 2023 Jan 10;120(2):e2216216120. doi: 10.1073/pnas.2216216120. Epub 2023 Jan 3. Proc Natl Acad Sci U S A. 2023. PMID: 36595701 Free PMC article.
References
-
- Biswas S. B., Kornberg A., 1984. Nucleoside triphosphate binding to DNA polymerase III holoenzyme of Escherichia coli. A direct photoaffinity labeling study. J. Biol. Chem. 259: 7990–7993. - PubMed
-
- Bloom L. B., Chen X., Fygenson D. K., Turner J., O’Donnell M., et al. , 1997. Fidelity of Escherichia coli DNA polymerase III holoenzyme. The effects of beta, gamma complex processivity proteins and epsilon proofreading exonuclease on nucleotide misincorporation efficiencies. J. Biol. Chem. 272: 27919–27930. 10.1074/jbc.272.44.27919 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

