Strain-level typing and identification of bacteria - a novel approach for SERS active plasmonic nanostructures
- PMID: 29907950
- PMCID: PMC6061775
- DOI: 10.1007/s00216-018-1153-0
Strain-level typing and identification of bacteria - a novel approach for SERS active plasmonic nanostructures
Abstract
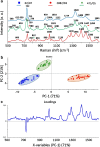
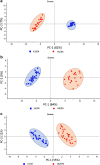
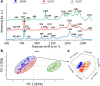
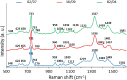
One of the potential applications of surface-enhanced Raman spectroscopy (SERS) is the detection of biological compounds and microorganisms. Here we demonstrate that SERS coupled with principal component analysis (PCA) serves as a perfect method for determining the taxonomic affiliation of bacteria at the strain level. We demonstrate for the first time that it is possible to distinguish different genoserogroups within a single species, Listeria monocytogenes, which is one of the most virulent foodborne pathogens and in some cases contact with which may be fatal. We also postulate that it is possible to detect additional proteins in the L. monocytogenes cell envelope, which provide resistance to benzalkonium chloride and cadmium. A better understanding of this infectious agent could help in selecting the appropriate pharmaceutical product for enhanced treatment. Graphical abstract ᅟ.
Keywords: Bacteria detection; Bacteria identification; BcrB; BcrC; CadA1; CadA2; Listeria monocytogenes; PCA; SERS; Strain level discrimination; bcrABC.
Conflict of interest statement
The authors declare that they have no conflict of interests.
Figures



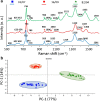
References
-
- Kneipp K, Wang Y, Kneipp H, Perelman LT, Itzkan I, Dasari RR, Feld MS. Single molecule detection using surface-enhanced raman scattering (SERS) Phys Rev Lett. 1997;78:1667–1670. doi: 10.1103/PhysRevLett.78.1667. - DOI
-
- Smith E, Dent G. Surface-enhanced Raman scattering and surface-enhanced resonance raman scattering. Modern Raman Spectroscopy – A Practical Approach. Hoboken: Wiley; 2005.
-
- Sägmüller B, Schwarze B, Brehm G, Trachta G, Schneider S. Identification of illicit drugs by a combination of liquid chromatography and surface-enhanced Raman scattering spectroscopy. J Mol Struct. 2003;661–662:279–290. doi: 10.1016/S0022-2860(03)00507-6. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

