Monitoring 15N Chemical Shifts During Protein Folding by Pressure-Jump NMR
- PMID: 29923716
- PMCID: PMC6119464
- DOI: 10.1021/jacs.8b04833
Monitoring 15N Chemical Shifts During Protein Folding by Pressure-Jump NMR
Abstract
Pressure-jump hardware permits direct observation of protein NMR spectra during a cyclically repeated protein folding process. For a two-state folding protein, the change in resonance frequency will occur nearly instantaneously when the protein clears the transition state barrier, resulting in a monoexponential change of the ensemble-averaged chemical shift. However, protein folding pathways can be more complex and contain metastable intermediates. With a pseudo-3D NMR experiment that utilizes stroboscopic observation, we measure the ensemble-averaged chemical shifts, including those of exchange-broadened intermediates, during the folding process. Such measurements for a pressure-sensitized mutant of ubiquitin show an on-pathway kinetic intermediate whose 15N chemical shifts differ most from the natively folded protein for strands β5, its preceding turn, and the two strands that pair with β5 in the native structure.
Figures
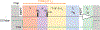

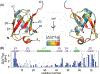
References
-
- Koide S; Dyson HJ; Wright PE, Characterization of a folding intermediate of apoplastocyanin trapped by proline isomerization. Biochemistry 1993, 32, 12299–12310. - PubMed
-
- Balbach J; Forge V; Vannuland NAJ; Winder SL; Hore PJ; Dobson CM, Following protein folding in real-time using NMR spectroscopy. Nat. Struct. Biol 1995, 2, 865–870. - PubMed
-
- Roche J; Dellarole M; Caro JA; Norberto DR; Garcia AE; Garcia-Moreno B; Roumestand C; Royer CA, Effect of Internal Cavities on Folding Rates and Routes Revealed by Real-Time Pressure-Jump NMR Spectroscopy. J. Am. Chem. Soc 2013, 135, 14610–14618. - PubMed
-
- Schlepckow K; Wirmer J; Bachmann A; Kiefhaber T; Schwalbe H, Conserved folding pathways of alpha-lactalbumin and lysozyme revealed by kinetic CD, fluorescence, NMR, and interrupted refolding experiments. J. Mol. Biol 2008, 378, 686–698. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

