Unsupervised clustering and epigenetic classification of single cells
- PMID: 29925875
- PMCID: PMC6010417
- DOI: 10.1038/s41467-018-04629-3
Unsupervised clustering and epigenetic classification of single cells
Abstract
Characterizing epigenetic heterogeneity at the cellular level is a critical problem in the modern genomics era. Assays such as single cell ATAC-seq (scATAC-seq) offer an opportunity to interrogate cellular level epigenetic heterogeneity through patterns of variability in open chromatin. However, these assays exhibit technical variability that complicates clear classification and cell type identification in heterogeneous populations. We present scABC, an R package for the unsupervised clustering of single-cell epigenetic data, to classify scATAC-seq data and discover regions of open chromatin specific to cell identity.
Conflict of interest statement
The authors declare no competing interests.
Figures
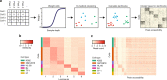

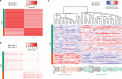
References
-
- Studer M. WeightedCluster library manual: a practical guide to creating typologies of trajectories in the social sciences with R. LIVES Work. Pap. 2013;24:1–34.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

