Genomewide association analyses of fitness traits in captive-reared Chinook salmon: Applications in evaluating conservation strategies
- PMID: 29928295
- PMCID: PMC5999212
- DOI: 10.1111/eva.12599
Genomewide association analyses of fitness traits in captive-reared Chinook salmon: Applications in evaluating conservation strategies
Abstract
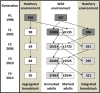
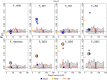
A novel application of genomewide association analyses is to use trait-associated loci to monitor the effects of conservation strategies on potentially adaptive genetic variation. Comparisons of fitness between captive- and wild-origin individuals, for example, do not reveal how captive rearing affects genetic variation underlying fitness traits or which traits are most susceptible to domestication selection. Here, we used data collected across four generations to identify loci associated with six traits in adult Chinook salmon (Oncorhynchus tshawytscha) and then determined how two alternative management approaches for captive rearing affected variation at these loci. Loci associated with date of return to freshwater spawning grounds (return timing), length and weight at return, age at maturity, spawn timing, and daily growth coefficient were identified using 9108 restriction site-associated markers and random forest, an approach suitable for polygenic traits. Mapping of trait-associated loci, gene annotations, and integration of results across multiple studies revealed candidate regions involved in several fitness-related traits. Genotypes at trait-associated loci were then compared between two hatchery populations that were derived from the same source but are now managed as separate lines, one integrated with and one segregated from the wild population. While no broad-scale change was detected across four generations, there were numerous regions where trait-associated loci overlapped with signatures of adaptive divergence previously identified in the two lines. Many regions, primarily with loci linked to return and spawn timing, were either unique to or more divergent in the segregated line, suggesting that these traits may be responding to domestication selection. This study is one of the first to utilize genomic approaches to demonstrate the effectiveness of a conservation strategy, managed gene flow, on trait-associated-and potentially adaptive-loci. The results will promote the development of trait-specific tools to better monitor genetic change in captive and wild populations.
Keywords: captive rearing; conservation; domestication selection; genomewide association analysis; managed gene flow; random forest.
Figures

Similar articles
-
Genomic and phenotypic effects of inbreeding across two different hatchery management regimes in Chinook salmon.Mol Ecol. 2020 Feb;29(4):658-672. doi: 10.1111/mec.15356. Epub 2020 Feb 14. Mol Ecol. 2020. PMID: 31957935
-
Effectiveness of managed gene flow in reducing genetic divergence associated with captive breeding.Evol Appl. 2015 Dec;8(10):956-71. doi: 10.1111/eva.12331. Epub 2015 Nov 4. Evol Appl. 2015. PMID: 26640521 Free PMC article.
-
Integration of Random Forest with population-based outlier analyses provides insight on the genomic basis and evolution of run timing in Chinook salmon (Oncorhynchus tshawytscha).Mol Ecol. 2015 Jun;24(11):2729-46. doi: 10.1111/mec.13211. Mol Ecol. 2015. PMID: 25913096
-
Evolution of chinook salmon (Oncorhynchus tshawytscha) populations in New Zealand: pattern, rate, and process.Genetica. 2001;112-113:493-513. Genetica. 2001. PMID: 11838785 Review.
-
On the reproductive success of early-generation hatchery fish in the wild.Evol Appl. 2014 Sep;7(8):883-96. doi: 10.1111/eva.12183. Epub 2014 Jul 9. Evol Appl. 2014. PMID: 25469167 Free PMC article. Review.
Cited by
-
Salmon hatchery strays can demographically boost wild populations at the cost of diversity: quantitative genetic modelling of Alaska pink salmon.R Soc Open Sci. 2024 Jul 3;11(7):240455. doi: 10.1098/rsos.240455. eCollection 2024 Jul. R Soc Open Sci. 2024. PMID: 39076353 Free PMC article.
-
A single generation in the wild increases fitness for descendants of hatchery-origin Chinook salmon (Oncorhynchus tshawytscha).Evol Appl. 2024 Apr 11;17(4):e13678. doi: 10.1111/eva.13678. eCollection 2024 Apr. Evol Appl. 2024. PMID: 38617826 Free PMC article.
-
Power of a dual-use SNP panel for pedigree reconstruction and population assignment.Ecol Evol. 2020 Aug 10;10(17):9522-9531. doi: 10.1002/ece3.6645. eCollection 2020 Sep. Ecol Evol. 2020. PMID: 32953080 Free PMC article.
-
Long-term evaluation of fitness and demographic effects of a Chinook Salmon supplementation program.Evol Appl. 2018 Nov 15;12(3):456-469. doi: 10.1111/eva.12725. eCollection 2019 Mar. Evol Appl. 2018. PMID: 30828367 Free PMC article.
-
Y-chromosome haplotypes are associated with variation in size and age at maturity in male Chinook salmon.Evol Appl. 2020 Aug 28;13(10):2791-2806. doi: 10.1111/eva.13084. eCollection 2020 Dec. Evol Appl. 2020. PMID: 33294023 Free PMC article.
References
-
- Akiduki, S. , & Ikemoto, M. J. (2008). Modulation of the neural glutamate transporter EAAC1 by the addicsin‐interacting protein ARL6IP1. Journal of Biological Chemistry, 283, 31323–31332. https://doi.org/10.1074/jbc.M801570200 - DOI - PubMed
-
- Allendorf, F. W. , & Thorgaard, G. H. (1984). Tetraploidy and the evolution of salmonid fishes In Turner B. J. (Ed.), Evolutionary genetics of fishes. New York, NY: Plenum Press.
-
- Altschul, S. F. , Gish, W. , Miller, W. , Myers, E. W. , & Lipman, D. J. (1990). Basic local alignment search tool. Journal of Molecular Biology, 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2 - DOI - PubMed
-
- Aykanat, T. , Lindqvist, M. , Pritchard, V. L. , & Primmer, C. R. (2016). From population genomics to conservation and management: A workflow for targeted analysis of markers identified using genome‐wide approaches in Atlantic salmon Salmo salar . Journal of Fish Biology, 89, 2658–2679. https://doi.org/10.1111/jfb.13149 - DOI - PubMed
-
- Baird, N. A. , Etter, P. D. , Atwood, T. S. , Currey, M. C. , Shiver, A. L. , Lewis, Z. A. , & Johnson, E. A. (2008). Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS One, 3, e3376 https://doi.org/10.1371/journal.pone.0003376 - DOI - PMC - PubMed
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources

