Impact of nutrition and rotavirus infection on the infant gut microbiota in a humanized pig model
- PMID: 29929472
- PMCID: PMC6013989
- DOI: 10.1186/s12876-018-0810-2
Impact of nutrition and rotavirus infection on the infant gut microbiota in a humanized pig model
Abstract
Background: Human rotavirus (HRV) is a major cause of viral gastroenteritis in infants; particularly in developing countries where malnutrition is prevalent. Malnutrition perturbs the infant gut microbiota leading to sub-optimal functioning of the immune system and further predisposing infants to enteric infections. Therefore, we hypothesized that malnutrition exacerbates rotavirus disease severity in infants.
Methods: In the present study, we used a neonatal germ free (GF) piglets transplanted with a two-month-old human infant's fecal microbiota (HIFM) on protein deficient and sufficient diets. We report the effects of malnourishment on the HRV infection and the HIFM pig microbiota in feces, intestinal and systemic tissues, using MiSeq 16S gene sequencing (V4-V5 region).
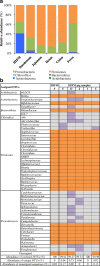
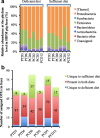
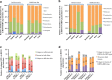
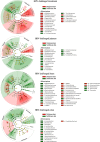
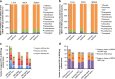
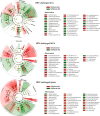
Results: Microbiota analysis indicated that the HIFM transplantation resulted in a microbial composition in pigs similar to that of the original infant feces. This model was then used to understand the interconnections between microbiota diversity, diet, and HRV infection. Post HRV infection, HIFM pigs on the deficient diet had lower body weights, developed more severe diarrhea and increased virus shedding compared to HIFM pigs on sufficient diet. However, HRV induced diarrhea and shedding was more pronounced in non-colonized GF pigs compared to HIFM pigs on either sufficient or deficient diet, suggesting that the microbiota alone moderated HRV infection. HRV infected pigs on sufficient diet showed increased microbiota diversity in intestinal tissues; whereas, greater diversity was observed in systemic tissues of HRV infected pigs fed with deficient diet.
Conclusions: These results suggest that proper nourishment improves the microbiota quality in the intestines, alleviates HRV disease and lower probability of systemic translocation of potential opportunistic pathogens/pathobionts. In conclusion, our findings further support the role for microbiota and proper nutrition in limiting enteric diseases.
Keywords: Humanized pig; Malnutrition; Microbiota; Protein diet; Rotavirus.
Conflict of interest statement
Ethics approval
All animal experiments were approved and performed in accordance to the Institutional Animal Care and Use Committee of The Ohio State University (Protocol #2010A00000088). Fecal sample collection from infant was conducted in accordance with the approved Ohio State University Institutional Review Board protocol (protocol #2016H0276). A written consent for publication was obtained from infant’s parent.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



References
-
- Mountzouris KC, McCartney AL, Gibson GR. Intestinal microflora of human infants and current trends for its nutritional modulation. Br J Nutr. 2002;87:405–420. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

