Correcting for targeted and control agent signal differences in paired-agent molecular imaging of cancer cell-surface receptors
- PMID: 29931837
- PMCID: PMC6013418
- DOI: 10.1117/1.JBO.23.6.066004
Correcting for targeted and control agent signal differences in paired-agent molecular imaging of cancer cell-surface receptors
Abstract
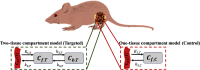
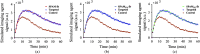
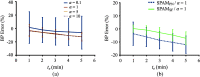
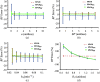
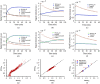
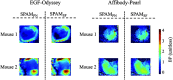
Paired-agent kinetic modeling protocols provide one means of estimating cancer cell-surface receptors with in vivo molecular imaging. The protocols employ the coadministration of a control imaging agent with one or more targeted imaging agent to account for the nonspecific uptake and retention of the targeted agent. These methods require the targeted and control agent data be converted to equivalent units of concentration, typically requiring specialized equipment and calibration, and/or complex algorithms that raise the barrier to adoption. This work evaluates a kinetic model capable of correcting for targeted and control agent signal differences. This approach was compared with an existing simplified paired-agent model (SPAM), and modified SPAM that accounts for signal differences by early time point normalization of targeted and control signals (SPAMPN). The scaling factor model (SPAMSF) outperformed both SPAM and SPAMPN in terms of accuracy and precision when the scale differences between targeted and imaging agent signals (α) were not equal to 1, and it matched the performance of SPAM for α = 1. This model could have wide-reaching implications for quantitative cancer receptor imaging using any imaging modalities, or combinations of imaging modalities, capable of concurrent detection of at least two distinct imaging agents (e.g., SPECT, optical, and PET/MR).
Keywords: kinetic modeling; optical tissue properties; paired-agent imaging.
(2018) COPYRIGHT Society of Photo-Optical Instrumentation Engineers (SPIE).
Figures
Similar articles
-
Generalized paired-agent kinetic model for in vivo quantification of cancer cell-surface receptors under receptor saturation conditions.Phys Med Biol. 2017 Jan 21;62(2):394-414. doi: 10.1088/1361-6560/62/2/394. Epub 2016 Dec 20. Phys Med Biol. 2017. PMID: 27997381 Free PMC article.
-
Quantifying cancer cell receptors with paired-agent fluorescent imaging: a novel method to account for tissue optical property effects.Proc SPIE Int Soc Opt Eng. 2018 Jan-Feb;10497:1049723. doi: 10.1117/12.2290631. Epub 2018 Feb 20. Proc SPIE Int Soc Opt Eng. 2018. PMID: 30220772 Free PMC article.
-
Accounting for pharmacokinetic differences in dual-tracer receptor density imaging.Phys Med Biol. 2014 May 21;59(10):2341-51. doi: 10.1088/0031-9155/59/10/2341. Epub 2014 Apr 17. Phys Med Biol. 2014. PMID: 24743262 Free PMC article.
-
Molecular imaging of breast cancer.Breast. 2009 Oct;18 Suppl 3:S66-73. doi: 10.1016/S0960-9776(09)70276-0. Breast. 2009. PMID: 19914546 Review.
-
Targeted nuclear imaging of breast cancer: status of radiotracer development and clinical applications.Cancer Biother Radiopharm. 2012 Mar;27(2):105-12. doi: 10.1089/cbr.2011.1025. Epub 2011 Aug 30. Cancer Biother Radiopharm. 2012. PMID: 21877909 Review.
Cited by
-
Noninvasive quantification of target availability during therapy using paired-agent fluorescence tomography.Theranostics. 2020 Sep 14;10(24):11230-11243. doi: 10.7150/thno.45273. eCollection 2020. Theranostics. 2020. PMID: 33042280 Free PMC article.
-
Dual-channel pulse-dye densitometry can enable correction of fluorescent targeted and control agent plasma input function differences for quantitative paired-agent molecular imaging: a simulation study.J Biomed Opt. 2025 Apr;30(4):046001. doi: 10.1117/1.JBO.30.4.046001. Epub 2025 Mar 29. J Biomed Opt. 2025. PMID: 40161250 Free PMC article.
-
Prediction of optimal contrast times post-imaging agent administration to inform personalized fluorescence-guided surgery.J Biomed Opt. 2020 Nov;25(11):116005. doi: 10.1117/1.JBO.25.11.116005. J Biomed Opt. 2020. PMID: 33200596 Free PMC article.
References
-
- Assaraf Y. G., Leamon C. P., Reddy J. A., “The folate receptor as a rational therapeutic target for personalized cancer treatment,” Drug Resist. Updat. 17(4), 89–95 (2014).https://doi.org/10.1016/j.drup.2014.10.002 - DOI - PubMed
-
- Henry K. E., Ulaner G. A., Lewis J. S., “Human epidermal growth factor receptor 2-targeted PET/single-photon emission computed tomography imaging of breast cancer: noninvasive measurement of a biomarker integral to tumor treatment and prognosis,” PET Clin. 12(3), 269–288 (2017).https://doi.org/10.1016/j.cpet.2017.02.001 - DOI - PMC - PubMed
-
- Clark P. M., et al. , “Harnessing preclinical molecular imaging to inform advances in personalized cancer medicine,” J. Nucl. Med. 58(5), 689–696 (2017).JNMEAQhttps://doi.org/10.2967/jnumed.116.181693 - DOI - PubMed
-
- Furnari F. B., et al. , “Heterogeneity of epidermal growth factor receptor signalling networks in glioblastoma,” Nat. Rev. Cancer 15(5), 302–310 (2015).NRCAC4https://doi.org/10.1038/nrc3918 - DOI - PMC - PubMed
-
- Gibbs-Strauss S. L., et al. , “Detecting epidermal growth factor receptor tumor activity in vivo during cetuximab therapy of murine gliomas,” Acad. Radiol. 17(1), 7–17 (2010).https://doi.org/10.1016/j.acra.2009.07.027 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

