Detecting Succinylation sites from protein sequences using ensemble support vector machine
- PMID: 29940836
- PMCID: PMC6016146
- DOI: 10.1186/s12859-018-2249-4
Detecting Succinylation sites from protein sequences using ensemble support vector machine
Abstract
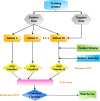
Background: Lysine succinylation is a new kind of post-translational modification which plays a key role in protein conformation regulation and cellular function control. To understand the mechanism of succinylation profoundly, it is necessary to identify succinylation sites in proteins accurately. However, traditional methods, experimental approaches, are labor-intensive and time-consuming. Computational prediction methods have been proposed recent years, and they are popular because of their convenience and high speed. In this study, we developed a new method to predict succinylation sites in protein combining multiple features, including amino acid composition, binary encoding, physicochemical property and grey pseudo amino acid composition, with a feature selection scheme (information gain). And then, it was trained using SVM (Support Vector Machine) and an ensemble learning algorithm.
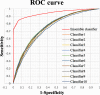
Results: The performance of this method was measured with an accuracy of 89.14% and a MCC (Matthew Correlation Coefficient) of 0.79 using 10-fold cross validation on training dataset and an accuracy of 84.5% and a MCC of 0.2 on independent dataset.
Conclusions: The conclusions made from this study can help to understand more of the succinylation mechanism. These results suggest that our method was very promising for predicting succinylation sites. The source code and data of this paper are freely available at https://github.com/ningq669/PSuccE .
Keywords: Ensemble learning algorithm; Grey pseudo amino acid composition; Information gain; Multiple features; Predict succinylation sites; SVM.
Conflict of interest statement
Ethics approval and consent to participate
All authors approval and consent to participate.
Consent for publication
All authors read and consent to publish the manuscript.
Competing interests
The authors declare that they no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures

Similar articles
-
Large-Scale Assessment of Bioinformatics Tools for Lysine Succinylation Sites.Cells. 2019 Jan 28;8(2):95. doi: 10.3390/cells8020095. Cells. 2019. PMID: 30696115 Free PMC article. Review.
-
Accurate in silico identification of protein succinylation sites using an iterative semi-supervised learning technique.J Theor Biol. 2015 Jun 7;374:60-5. doi: 10.1016/j.jtbi.2015.03.029. Epub 2015 Apr 2. J Theor Biol. 2015. PMID: 25843215
-
SSKM_Succ: A Novel Succinylation Sites Prediction Method Incorporating K-Means Clustering With a New Semi-Supervised Learning Algorithm.IEEE/ACM Trans Comput Biol Bioinform. 2022 Jan-Feb;19(1):643-652. doi: 10.1109/TCBB.2020.3006144. Epub 2022 Feb 3. IEEE/ACM Trans Comput Biol Bioinform. 2022. PMID: 32750881
-
SuccinSite: a computational tool for the prediction of protein succinylation sites by exploiting the amino acid patterns and properties.Mol Biosyst. 2016 Mar;12(3):786-95. doi: 10.1039/c5mb00853k. Epub 2016 Jan 7. Mol Biosyst. 2016. PMID: 26739209
-
A Comprehensive Comparative Review of Protein Sequence-Based Computational Prediction Models of Lysine Succinylation Sites.Curr Protein Pept Sci. 2022;23(11):744-756. doi: 10.2174/1389203723666220628121817. Curr Protein Pept Sci. 2022. PMID: 35762552 Review.
Cited by
-
Improving protein succinylation sites prediction using embeddings from protein language model.Sci Rep. 2022 Oct 8;12(1):16933. doi: 10.1038/s41598-022-21366-2. Sci Rep. 2022. PMID: 36209286 Free PMC article.
-
Large-Scale Assessment of Bioinformatics Tools for Lysine Succinylation Sites.Cells. 2019 Jan 28;8(2):95. doi: 10.3390/cells8020095. Cells. 2019. PMID: 30696115 Free PMC article. Review.
-
Succinylation Site Prediction Based on Protein Sequences Using the IFS-LightGBM (BO) Model.Comput Math Methods Med. 2020 Nov 10;2020:8858489. doi: 10.1155/2020/8858489. eCollection 2020. Comput Math Methods Med. 2020. PMID: 33224267 Free PMC article.
-
LMPTMSite: A Platform for PTM Site Prediction in Proteins Leveraging Transformer-Based Protein Language Models.Methods Mol Biol. 2025;2867:261-297. doi: 10.1007/978-1-0716-4196-5_16. Methods Mol Biol. 2025. PMID: 39576587
-
Prediction and analysis of multiple protein lysine modified sites based on conditional wasserstein generative adversarial networks.BMC Bioinformatics. 2021 Mar 31;22(1):171. doi: 10.1186/s12859-021-04101-y. BMC Bioinformatics. 2021. PMID: 33789579 Free PMC article.
References
-
- Tan M, Peng C, Anderson K, Chhoy P, Xie Z, Dai L, Park J, Chen Y, Huang H, Zhang Y, Ro J, Wagner GR, Green MF, Madsen AS, Schmiesing J, Peterson BS, Xu G, Ilkayeva OR, Muehlbauer MJ, Braulke T, Mühlhausen C, Backos DS, Olsen CA, McGuire PJ, Pletcher SD, Lombard DB, Hirschey MD, Zhao Y. Lysine Glutarylation is a protein posttranslational modification regulated by SIRT5 [J] Cell Metab. 2014;19(4):605–617. doi: 10.1016/j.cmet.2014.03.014. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

