Creating Lineage Trajectory Maps Via Integration of Single-Cell RNA-Sequencing and Lineage Tracing: Integrating transgenic lineage tracing and single-cell RNA-sequencing is a robust approach for mapping developmental lineage trajectories and cell fate changes
- PMID: 29944188
- PMCID: PMC6161781
- DOI: 10.1002/bies.201800056
Creating Lineage Trajectory Maps Via Integration of Single-Cell RNA-Sequencing and Lineage Tracing: Integrating transgenic lineage tracing and single-cell RNA-sequencing is a robust approach for mapping developmental lineage trajectories and cell fate changes
Abstract
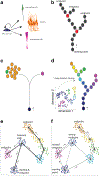
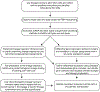
Mapping the paths that stem and progenitor cells take en route to differentiate and elucidating the underlying molecular controls are key goals in developmental and stem cell biology. However, with population level analyses it is difficult - if not impossible - to define the transition states and lineage trajectory branch points within complex developmental lineages. Single-cell RNA-sequencing analysis can discriminate heterogeneity in a population of cells and even identify rare or transient intermediates. In this review, we propose that using these data, one can infer the lineage trajectories of individual stem cells and identify putative branch points. Clonal lineage tracing of stem cells allows one to define the outcome of differentiation. Integrating these single cell-based approaches provides a robust strategy for establishing and testing models of how an individual stem cell changes through time to differentiate and self-renew.
Keywords: cell fate; lineage; scRNA-seq; single-cell RNA-sequencing; stem cells.
© 2018 WILEY Periodicals, Inc.
Conflict of interest statement
Conflict of Interest
The authors declare no conflict of interest.
Figures


Similar articles
-
LineageOT is a unified framework for lineage tracing and trajectory inference.Nat Commun. 2021 Aug 16;12(1):4940. doi: 10.1038/s41467-021-25133-1. Nat Commun. 2021. PMID: 34400634 Free PMC article.
-
GEMLI: Gene Expression Memory-Based Lineage Inference from Single-Cell RNA-Sequencing Datasets.Methods Mol Biol. 2025;2886:375-400. doi: 10.1007/978-1-0716-4310-5_19. Methods Mol Biol. 2025. PMID: 39745651
-
Single-cell analysis of mixed-lineage states leading to a binary cell fate choice.Nature. 2016 Sep 29;537(7622):698-702. doi: 10.1038/nature19348. Epub 2016 Aug 31. Nature. 2016. PMID: 27580035 Free PMC article.
-
Next-Generation Lineage Tracing and Fate Mapping to Interrogate Development.Dev Cell. 2021 Jan 11;56(1):7-21. doi: 10.1016/j.devcel.2020.10.021. Epub 2020 Nov 19. Dev Cell. 2021. PMID: 33217333 Review.
-
Constructing cell lineages from single-cell transcriptomes.Mol Aspects Med. 2018 Feb;59:95-113. doi: 10.1016/j.mam.2017.10.004. Epub 2017 Nov 26. Mol Aspects Med. 2018. PMID: 29107741 Review.
Cited by
-
Inferring changes in histone modification during cell differentiation by ancestral state estimation based on phylogenetic trees of cell types: Human hematopoiesis as a model case.Gene X. 2019 May 31;3:100021. doi: 10.1016/j.gene.2019.100021. eCollection 2019 Sep. Gene X. 2019. PMID: 32550550 Free PMC article.
-
Identifying gene expression from single cells to single genes.Methods Cell Biol. 2019;151:127-158. doi: 10.1016/bs.mcb.2018.11.018. Epub 2019 Jan 2. Methods Cell Biol. 2019. PMID: 30948004 Free PMC article. Review.
-
A functional cellular framework for sex and estrous cycle-dependent gene expression and behavior.Cell. 2022 Feb 17;185(4):654-671.e22. doi: 10.1016/j.cell.2021.12.031. Epub 2022 Jan 21. Cell. 2022. PMID: 35065713 Free PMC article.
-
LineageOT is a unified framework for lineage tracing and trajectory inference.Nat Commun. 2021 Aug 16;12(1):4940. doi: 10.1038/s41467-021-25133-1. Nat Commun. 2021. PMID: 34400634 Free PMC article.
-
The roles of microRNAs in mouse development.Nat Rev Genet. 2021 May;22(5):307-323. doi: 10.1038/s41576-020-00309-5. Epub 2021 Jan 15. Nat Rev Genet. 2021. PMID: 33452500 Review.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

