Convergent evolution of integration site selection upstream of tRNA genes by yeast and amoeba retrotransposons
- PMID: 29945249
- PMCID: PMC6101501
- DOI: 10.1093/nar/gky582
Convergent evolution of integration site selection upstream of tRNA genes by yeast and amoeba retrotransposons
Abstract
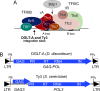
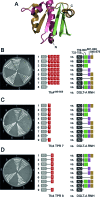
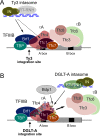
Transposable elements amplify in genomes as selfish DNA elements and challenge host fitness because their intrinsic integration steps during mobilization can compromise genome integrity. In gene-dense genomes, transposable elements are notably under selection to avoid insertional mutagenesis of host protein-coding genes. We describe an example of convergent evolution in the distantly related amoebozoan Dictyostelium discoideum and the yeast Saccharomyces cerevisiae, in which the D. discoideum retrotransposon DGLT-A and the yeast Ty3 element developed different mechanisms to facilitate position-specific integration at similar sites upstream of tRNA genes. Transcription of tRNA genes by RNA polymerase III requires the transcription factor complexes TFIIIB and TFIIIC. Whereas Ty3 recognizes tRNA genes mainly through interactions of its integrase with TFIIIB subunits, the DGLT-A-encoded ribonuclease H contacts TFIIIC subunit Tfc4 at an interface that covers tetratricopeptide repeats (TPRs) 7 and 8. A major function of this interface is to connect TFIIIC subcomplexes τA and τB and to facilitate TFIIIB assembly. During the initiation of tRNA gene transcription τB is displaced from τA, which transiently exposes the TPR 7/8 surface of Tfc4 on τA. We propose that the DGLT-A intasome uses this binding site to obtain access to genomic DNA for integration during tRNA gene transcription.
Figures




Similar articles
-
TFIIIC-independent in vitro transcription of yeast tRNA genes.J Mol Biol. 2000 Jun 9;299(3):601-13. doi: 10.1006/jmbi.2000.3783. J Mol Biol. 2000. PMID: 10835271
-
Protein interactions involved in tRNA gene-specific integration of Dictyostelium discoideum non-long terminal repeat retrotransposon TRE5-A.Mol Cell Biol. 2007 Dec;27(24):8492-501. doi: 10.1128/MCB.01173-07. Epub 2007 Oct 8. Mol Cell Biol. 2007. PMID: 17923679 Free PMC article.
-
Structural basis of Ty3 retrotransposon integration at RNA Polymerase III-transcribed genes.Nat Commun. 2021 Nov 30;12(1):6992. doi: 10.1038/s41467-021-27338-w. Nat Commun. 2021. PMID: 34848735 Free PMC article.
-
Regulation of tRNA synthesis by the general transcription factors of RNA polymerase III - TFIIIB and TFIIIC, and by the MAF1 protein.Biochim Biophys Acta Gene Regul Mech. 2018 Apr;1861(4):320-329. doi: 10.1016/j.bbagrm.2018.01.011. Epub 2018 Feb 6. Biochim Biophys Acta Gene Regul Mech. 2018. PMID: 29378333 Review.
-
Comparison of the RNA polymerase III transcription machinery in Schizosaccharomyces pombe, Saccharomyces cerevisiae and human.Nucleic Acids Res. 2001 Jul 1;29(13):2675-90. doi: 10.1093/nar/29.13.2675. Nucleic Acids Res. 2001. PMID: 11433012 Free PMC article. Review.
Cited by
-
A small targeting domain in Ty1 integrase is sufficient to direct retrotransposon integration upstream of tRNA genes.EMBO J. 2020 Sep 1;39(17):e104337. doi: 10.15252/embj.2019104337. Epub 2020 Jul 17. EMBO J. 2020. PMID: 32677087 Free PMC article.
References
-
- Orgel L.E., Crick F.H.C.. Selfish DNA: the ultimate parasite. Nature. 1980; 284:604–607. - PubMed
-
- Doolittle W.F., Sapienza C.. Selfish genes, the phenotype paradigm and genome evolution. Nature. 1980; 284:601–603. - PubMed
-
- Sultana T., Zamborlini A., Cristofari G., Lesage P.. Integration site selection by retroviruses and transposable elements in eukaryotes. Nat. Rev. Genet. 2017; 18:292–308. - PubMed
-
- Kim J.M., Vanguri S., Boeke J.D., Gabriel A., Voytas D.F.. Transposable elements and genome organization: a comprehensive survey of retrotransposons revealed by the complete Saccharomyces cerevisiae genome sequence. Genome Res. 1998; 8:464–478. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

