Conservation and divergence of YODA MAPKKK function in regulation of grass epidermal patterning
- PMID: 29945871
- PMCID: PMC6078329
- DOI: 10.1242/dev.165860
Conservation and divergence of YODA MAPKKK function in regulation of grass epidermal patterning
Abstract
All multicellular organisms must properly pattern cell types to generate functional tissues and organs. The organized and predictable cell lineages of the Brachypodium leaf enabled us to characterize the role of the MAPK kinase kinase gene BdYODA1 in regulating asymmetric cell divisions. We find that YODA genes promote normal stomatal spacing patterns in both Arabidopsis and Brachypodium, despite species-specific differences in those patterns. Using lineage tracing and cell fate markers, we show that, unexpectedly, patterning defects in bdyoda1 mutants do not arise from faulty physical asymmetry in cell divisions but rather from improper enforcement of alternative cellular fates after division. These cross-species comparisons allow us to refine our understanding of MAPK activities during plant asymmetric cell divisions.
Keywords: Asymmetric cell division; Brachypodium; Comparative development; MAPK pathway; Stomata.
© 2018. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures





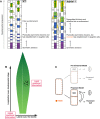
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

