POU2F3 is a master regulator of a tuft cell-like variant of small cell lung cancer
- PMID: 29945888
- PMCID: PMC6075037
- DOI: 10.1101/gad.314815.118
POU2F3 is a master regulator of a tuft cell-like variant of small cell lung cancer
Abstract
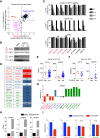
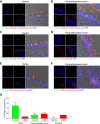
Small cell lung cancer (SCLC) is widely considered to be a tumor of pulmonary neuroendocrine cells; however, a variant form of this disease has been described that lacks neuroendocrine features. Here, we applied domain-focused CRISPR screening to human cancer cell lines to identify the transcription factor (TF) POU2F3 (POU class 2 homeobox 3; also known as SKN-1a/OCT-11) as a powerful dependency in a subset of SCLC lines. An analysis of human SCLC specimens revealed that POU2F3 is expressed exclusively in variant SCLC tumors that lack expression of neuroendocrine markers and instead express markers of a chemosensory lineage known as tuft cells. Using chromatin- and RNA-profiling experiments, we provide evidence that POU2F3 is a master regulator of tuft cell identity in a variant form of SCLC. Moreover, we show that most SCLC tumors can be classified into one of three lineages based on the expression of POU2F3, ASCL1, or NEUROD1. Our CRISPR screens exposed other unique dependencies in POU2F3-expressing SCLC lines, including the lineage TFs SOX9 and ASCL2 and the receptor tyrosine kinase IGF1R (insulin-like growth factor 1 receptor). These data reveal POU2F3 as a cell identity determinant and a dependency in a tuft cell-like variant of SCLC, which may reflect a previously unrecognized cell of origin or a trans-differentiation event in this disease.
Keywords: POU2F3; enhancer; master regulator; small cell lung cancer; tuft cell.
© 2018 Huang et al.; Published by Cold Spring Harbor Laboratory Press.
Figures



References
-
- Andersen B, Weinberg WC, Rennekampff O, McEvilly RJ, Bermingham JR Jr, Hooshmand F, Vasilyev V, Hansbrough JF, Pittelkow MR, Yuspa SH, et al. 1997. Functions of the POU domain genes Skn-1a/i and Tst-1/Oct-6/SCIP in epidermal differentiation. Genes Dev 11: 1873–1884. - PubMed
-
- Bezencon C, Furholz A, Raymond F, Mansourian R, Metairon S, Le Coutre J, Damak S. 2008. Murine intestinal cells expressing Trpm5 are mostly brush cells and express markers of neuronal and inflammatory cells. J Comp Neurol 509: 514–525. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous
