Mapping of ribosomal 23S ribosomal RNA modifications in Clostridium sporogenes
- PMID: 29947286
- PMCID: PMC6161674
- DOI: 10.1080/15476286.2018.1486662
Mapping of ribosomal 23S ribosomal RNA modifications in Clostridium sporogenes
Abstract
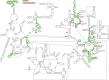
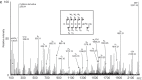
All organisms contain RNA modifications in their ribosomal RNA (rRNA), but the importance, positions and exact function of these are still not fully elucidated. Various functions such as stabilizing structures, controlling ribosome assembly and facilitating interactions have been suggested and in some cases substantiated. Bacterial rRNA contains much fewer modifications than eukaryotic rRNA. The rRNA modification patterns in bacteria differ from each other, but too few organisms have been mapped to draw general conclusions. This study maps 23S ribosomal RNA modifications in Clostridium sporogenes that can be characterized as a non-toxin producing Clostridium botulinum. Clostridia are able to sporulate and thereby survive harsh conditions, and are in general considered to be resilient to antibiotics. Selected regions of the 23S rRNA were investigated by mass spectrometry and by primer extension analysis to pinpoint modified sites and the nature of the modifications. Apparently, C. sporogenes 23S rRNA contains few modifications compared to other investigated bacteria. No modifications were identified in domain II and III of 23S rRNA. Three modifications were identified in domain IV, all of which have also been found in other organisms. Two unusual modifications were identified in domain V, methylated dihydrouridine at position U2449 and dihydrouridine at position U2500 (Escherichia coli numbering), in addition to four previously known modified positions. The enzymes responsible for the modifications were searched for in the C. sporogenes genome using BLAST with characterized enzymes as query. The search identified genes potentially coding for RNA modifying enzymes responsible for most of the found modifications.
Keywords: 23S RNA; RNA methylations; RNA modifications; dihydrouridine; mass spectroscopy; oh5C.
Figures



Similar articles
-
The archaeon Haloarcula marismortui has few modifications in the central parts of its 23S ribosomal RNA.J Mol Biol. 2005 May 6;348(3):563-73. doi: 10.1016/j.jmb.2005.03.009. J Mol Biol. 2005. PMID: 15826654
-
Posttranscriptional modifications in the A-loop of 23S rRNAs from selected archaea and eubacteria.RNA. 2002 Feb;8(2):202-13. doi: 10.1017/s1355838202013365. RNA. 2002. PMID: 11911366 Free PMC article.
-
Mutagenesis of the peptidyltransferase center of 23S rRNA: the invariant U2449 is dispensable.Nucleic Acids Res. 2001 Feb 1;29(3):710-5. doi: 10.1093/nar/29.3.710. Nucleic Acids Res. 2001. PMID: 11160893 Free PMC article.
-
Reprint of New opportunities for improved ribotyping of C. difficile clinical isolates by exploring their genomes.J Microbiol Methods. 2013 Dec;95(3):425-40. doi: 10.1016/j.mimet.2013.09.009. Epub 2013 Sep 16. J Microbiol Methods. 2013. PMID: 24050948 Review.
-
Naturally occurring modified ribonucleosides.Wiley Interdiscip Rev RNA. 2020 Sep;11(5):e1595. doi: 10.1002/wrna.1595. Epub 2020 Apr 16. Wiley Interdiscip Rev RNA. 2020. PMID: 32301288 Free PMC article. Review.
Cited by
-
Complete list of canonical post-transcriptional modifications in the Bacillus subtilis ribosome and their link to RbgA driven large subunit assembly.bioRxiv [Preprint]. 2024 May 11:2024.05.10.593627. doi: 10.1101/2024.05.10.593627. bioRxiv. 2024. Update in: Nucleic Acids Res. 2024 Oct 14;52(18):11203-11217. doi: 10.1093/nar/gkae626. PMID: 38765983 Free PMC article. Updated. Preprint.
-
Complete list of canonical post-transcriptional modifications in the Bacillus subtilis ribosome and their link to RbgA driven large subunit assembly.Nucleic Acids Res. 2024 Oct 14;52(18):11203-11217. doi: 10.1093/nar/gkae626. Nucleic Acids Res. 2024. PMID: 39036956 Free PMC article.
-
TlyA is a 23S and 16S 2'-O-methylcytidine methyltransferase important for ribosome assembly in Bacillus subtilis.bioRxiv [Preprint]. 2025 Apr 25:2025.04.21.649808. doi: 10.1101/2025.04.21.649808. bioRxiv. 2025. PMID: 40406463 Free PMC article. Preprint.
-
Structural insight into the novel Thermus thermophilus SPOUT methyltransferase RlmR catalysing Um2552 formation in the 23S rRNA A-loop: a case of convergent evolution.Nucleic Acids Res. 2025 May 22;53(10):gkaf432. doi: 10.1093/nar/gkaf432. Nucleic Acids Res. 2025. PMID: 40444636 Free PMC article.
-
Dihydrouridine synthesis in tRNAs is under reductive evolution in Mollicutes.RNA Biol. 2021 Dec;18(12):2278-2289. doi: 10.1080/15476286.2021.1899653. Epub 2021 Mar 22. RNA Biol. 2021. PMID: 33685366 Free PMC article.
References
-
- Decatur WA, Fournier MJ.. rRNA modifications and ribosome function. Trends Biochem Sci. 2002. July;27(7):344–351. PubMed PMID: 12114023. - PubMed
-
- Sergeeva OV, Bogdanov AA, Sergiev PV. What do we know about ribosomal RNA methylation in Escherichia coli? Biochimie. 2015. October;117:110–118. PubMed PMID: 25511423. - PubMed
-
- Fischer N, Neumann P, Konevega AL, et al. Structure of the E. coli ribosome-EF-Tu complex at <3 A resolution by Cs-corrected cryo-EM. Nature. 2015. April 23;520(7548):567–570. PubMed PMID: 25707802. - PubMed
-
- Mengel-Jorgensen J, Jensen SS, Rasmussen A, et al. Modifications in Thermus thermophilus 23 S ribosomal RNA are centered in regions of RNA-RNA contact. J Biol Chem. 2006. August 4;281(31):22108–22117. PubMed PMID: 16731530. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
