The Bacteriophage T4 MotB Protein, a DNA-Binding Protein, Improves Phage Fitness
- PMID: 29949907
- PMCID: PMC6070864
- DOI: 10.3390/v10070343
The Bacteriophage T4 MotB Protein, a DNA-Binding Protein, Improves Phage Fitness
Abstract
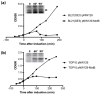
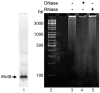
The lytic bacteriophage T4 employs multiple phage-encoded early proteins to takeover the Escherichia coli host. However, the functions of many of these proteins are not known. In this study, we have characterized the T4 early gene motB, located in a dispensable region of the T4 genome. We show that heterologous production of MotB is highly toxic to E. coli, resulting in cell death or growth arrest depending on the strain and that the presence of motB increases T4 burst size 2-fold. Previous work suggested that motB affects middle gene expression, but our transcriptome analyses of T4 motBam vs. T4 wt infections reveal that only a few late genes are mildly impaired at 5 min post-infection, and expression of early and middle genes is unaffected. We find that MotB is a DNA-binding protein that binds both unmodified host and T4 modified [(glucosylated, hydroxymethylated-5 cytosine, (GHme-C)] DNA with no detectable sequence specificity. Interestingly, MotB copurifies with the host histone-like proteins, H-NS and StpA, either directly or through cobinding to DNA. We show that H-NS also binds modified T4 DNA and speculate that MotB may alter how H-NS interacts with T4 DNA, host DNA, or both, thereby improving the growth of the phage.
Keywords: DNA-binding protein; H-NS; MotB; RNA-seq; bacteriophage T4; bacteriostatic; host takeover; transcriptome analysis.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

