The presence of genetic risk variants within PTPN2 and PTPN22 is associated with intestinal microbiota alterations in Swiss IBD cohort patients
- PMID: 29965986
- PMCID: PMC6028086
- DOI: 10.1371/journal.pone.0199664
The presence of genetic risk variants within PTPN2 and PTPN22 is associated with intestinal microbiota alterations in Swiss IBD cohort patients
Abstract
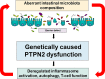
Background: Genetic risk factors, intestinal microbiota and a dysregulated immune system contribute to the pathogenesis of inflammatory bowel disease (IBD). We have previously demonstrated that dysfunction of protein tyrosine phosphatase non-receptor type 2 (PTPN2) and PTPN22 contributes to alterations of intestinal microbiota and the onset of chronic intestinal inflammation in vivo. Here, we investigated the influence of PTPN2 and PTPN22 gene variants on intestinal microbiota composition in IBD patients.
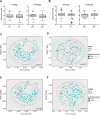
Methods: Bacterial DNA from mucosa-associated samples of 75 CD and 57 UC patients were sequenced using 16S rRNA sequencing approach. Microbial analysis, including alpha diversity, beta diversity and taxonomical analysis by comparing to PTPN2 (rs1893217) and PTPN22 (rs2476601) genotypes was performed in QIIME, the phyloseq R package and MaAsLin pipeline.
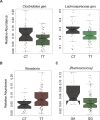
Results: In PTPN2 variant UC patients, we detected an increase in relative abundance of unassigned genera from Clostridiales and Lachnospiraceae families and reduction of Roseburia when compared to PTPN2 wild-type (WT) patients. Ruminoccocus was increased in PTPN22 variant UC patients. In CD patients with severe disease course, Faecalibacterium, Bilophila, Coprococcus, unclassified Erysipelotrichaeceae, unassigned genera from Clostridiales and Ruminococcaceae families were reduced and Bacteroides were increased in PTPN2 WT carriers, while Faecalibacterium, Bilophila, Coprococcus, and Erysipelotrichaeceae were reduced in PTPN22 WT patients when compared to patients with mild disease. In UC patients with severe disease, relative abundance of Lachnobacterium was reduced in PTPN2 and PTPN22 WT patients, Dorea was increased in samples from PTPN22 WT carriers and an unassigned genus from Ruminococcaceae gen. was increased in patients with PTPN2 variant genotype.
Conclusions: We identified that IBD-associated genetic risk variants, disease severity and the interaction of these factors are related to significant alterations in intestinal microbiota composition of IBD patients.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
The Clinical Relevance of the IBD-Associated Variation within the Risk Gene Locus Encoding Protein Tyrosine Phosphatase Non-Receptor Type 2 in Patients of the Swiss IBD Cohort.Digestion. 2016;93(3):182-92. doi: 10.1159/000444479. Epub 2016 Feb 27. Digestion. 2016. PMID: 26928573
-
Genotype-Phenotype Associations of the CD-Associated Single Nucleotide Polymorphism within the Gene Locus Encoding Protein Tyrosine Phosphatase Non-Receptor Type 22 in Patients of the Swiss IBD Cohort.PLoS One. 2016 Jul 28;11(7):e0160215. doi: 10.1371/journal.pone.0160215. eCollection 2016. PLoS One. 2016. PMID: 27467733 Free PMC article.
-
Polymorphisms in Protein Tyrosine Phosphatase Non-receptor Type 2 and 22 (PTPN2/22) Are Linked to Hyper-Proliferative T-Cells and Susceptibility to Mycobacteria in Rheumatoid Arthritis.Front Cell Infect Microbiol. 2018 Jan 25;8:11. doi: 10.3389/fcimb.2018.00011. eCollection 2018. Front Cell Infect Microbiol. 2018. PMID: 29423382 Free PMC article.
-
Protein tyrosine phosphatase non-receptor type 2 and inflammatory bowel disease.World J Gastroenterol. 2016 Jan 21;22(3):1034-44. doi: 10.3748/wjg.v22.i3.1034. World J Gastroenterol. 2016. PMID: 26811645 Free PMC article. Review.
-
Genetic Variations of PTPN2 and PTPN22: Role in the Pathogenesis of Type 1 Diabetes and Crohn's Disease.Front Cell Infect Microbiol. 2015 Dec 24;5:95. doi: 10.3389/fcimb.2015.00095. eCollection 2015. Front Cell Infect Microbiol. 2015. PMID: 26734582 Free PMC article. Review.
Cited by
-
Identification of a Novel Susceptibility Marker for SARS-CoV-2 Infection in Human Subjects and Risk Mitigation with a Clinically Approved JAK Inhibitor in Human/Mouse Cells.bioRxiv [Preprint]. 2020 Dec 9:2020.12.09.416586. doi: 10.1101/2020.12.09.416586. bioRxiv. 2020. PMID: 33330862 Free PMC article. Preprint.
-
Gut Microbiota Profiles Differ among Individuals Depending on Their Region of Origin: An Italian Pilot Study.Int J Environ Res Public Health. 2019 Oct 23;16(21):4065. doi: 10.3390/ijerph16214065. Int J Environ Res Public Health. 2019. PMID: 31652705 Free PMC article.
-
Cancer-Preventive Role of Bone Marrow-Derived Mesenchymal Stem Cells on Colitis-Associated Colorectal Cancer: Roles of Gut Microbiota Involved.Front Cell Dev Biol. 2021 Jun 4;9:642948. doi: 10.3389/fcell.2021.642948. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34150751 Free PMC article.
-
Autoimmune susceptibility gene PTPN2 is required for clearance of adherent-invasive Escherichia coli by integrating bacterial uptake and lysosomal defence.Gut. 2022 Jan;71(1):89-99. doi: 10.1136/gutjnl-2020-323636. Epub 2021 Feb 9. Gut. 2022. PMID: 33563644 Free PMC article.
-
PTPN2 Regulates Interactions Between Macrophages and Intestinal Epithelial Cells to Promote Intestinal Barrier Function.Gastroenterology. 2020 Nov;159(5):1763-1777.e14. doi: 10.1053/j.gastro.2020.07.004. Epub 2020 Jul 9. Gastroenterology. 2020. PMID: 32652144 Free PMC article.
References
-
- Hooper LV, Littman DR, Macpherson AJ. Interactions between the microbiota and the immune system. Science. 2012;336(6086):1268–73. doi: 10.1126/science.1223490 ; PubMed Central PMCID: PMC4420145. - DOI - PMC - PubMed
-
- Yilmaz B, Portugal S, Tran TM, Gozzelino R, Ramos S, Gomes J, et al. Gut microbiota elicits a protective immune response against malaria transmission. Cell. 2014;159(6):1277–89. doi: 10.1016/j.cell.2014.10.053 ; PubMed Central PMCID: PMC4261137. - DOI - PMC - PubMed
-
- Clemente JC, Ursell LK, Parfrey LW, Knight R. The impact of the gut microbiota on human health: an integrative view. Cell. 2012;148(6):1258–70. doi: 10.1016/j.cell.2012.01.035 ; PubMed Central PMCID: PMCPMC5050011. - DOI - PMC - PubMed
-
- Kaur N, Chen CC, Luther J, Kao JY. Intestinal dysbiosis in inflammatory bowel disease. Gut Microbes. 2011;2(4):211–6. doi: 10.4161/gmic.2.4.17863 . - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

