Manipulation of two regulatory genes for efficient production of chromomycins in Streptomyces reseiscleroticus
- PMID: 29977332
- PMCID: PMC5992853
- DOI: 10.1186/s13036-018-0103-x
Manipulation of two regulatory genes for efficient production of chromomycins in Streptomyces reseiscleroticus
Abstract
Background: Regulatory genes play critical roles in natural product biosynthetic pathways. Chromomycins are promising anticancer natural products from actinomycetes. This study is aimed to create an efficient strain for production of these molecules by manipulating the regulatory genes.
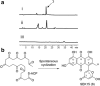
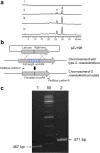
Results: A putative but silent chromomycin biosynthetic gene cluster was discovered in Streptomyces reseiscleroticus. Heterologous expression of the ketosynthase, chain length factor, and acyl carrier protein in Streptomyces lividans confirmed that they are responsible for the assembly of a decaketide. Two regulatory genes are present in this gene cluster, including SARP-type activator SrcmRI and PadR-like repressor SrcmRII. Either overexpression of SrcmRI or disruption of SrcmRII turned on the biosynthetic pathway of chromomycins. The production titers of chromomycin A3/A2 in R5 agar in these two strains reached 8.9 ± 1.2/13.2 ± 1.6 and 49.3 ± 4.3/53.3 ± 3.6 mg/L, respectively. An engineered strain was then constructed with both SrcmRII disruption and SrcmRI overexpression, which produced chromomycins A3 and A2 in R5 agar at 69.4 ± 7.6 and 81.7 ± 7.2 mg/L, respectively. Optimization of the culture conditions further increased the titers of chromomycins A3 and A2 respectively to 145.1 ± 15.3 and 158.3 ± 15.4 mg/L in liquid fermentation.
Conclusions: This work revealed the synergistic effect of manipulation of pathway repressor and activator genes in the engineering of a natural product biosynthetic pathway. The resulting engineered strain showed the highest production titers of chromomycins by a strain of Streptomyces, providing an efficient way to produce these pharmaceutically valuable molecules.
Keywords: Biosynthesis; Chromomycins; PadR-like transcription regulator; SARP regulator; Streptomyces reseiscleroticus.
Conflict of interest statement
Not applicable.The authors declare that they have no competing interests.Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



Similar articles
-
Identification and utility of FdmR1 as a Streptomyces antibiotic regulatory protein activator for fredericamycin production in Streptomyces griseus ATCC 49344 and heterologous hosts.J Bacteriol. 2008 Aug;190(16):5587-96. doi: 10.1128/JB.00592-08. Epub 2008 Jun 13. J Bacteriol. 2008. PMID: 18556785 Free PMC article.
-
Heterologous expression of pikromycin biosynthetic gene cluster using Streptomyces artificial chromosome system.Microb Cell Fact. 2017 May 31;16(1):96. doi: 10.1186/s12934-017-0708-7. Microb Cell Fact. 2017. PMID: 28569150 Free PMC article.
-
Isolation and identification of newly isolated antagonistic Streptomyces sp. strain AP19-2 producing chromomycins.J Microbiol. 2007 Dec;45(6):499-504. J Microbiol. 2007. PMID: 18176531
-
Antibacterial Activity of Chromomycins from a Marine-Derived Streptomyces microflavus.Mar Drugs. 2020 Oct 21;18(10):522. doi: 10.3390/md18100522. Mar Drugs. 2020. PMID: 33096696 Free PMC article.
-
Cloning and Heterologous Expression of a Large-sized Natural Product Biosynthetic Gene Cluster in Streptomyces Species.Front Microbiol. 2017 Mar 15;8:394. doi: 10.3389/fmicb.2017.00394. eCollection 2017. Front Microbiol. 2017. PMID: 28360891 Free PMC article. Review.
Cited by
-
Structural and functional characterization of AfsR, an SARP family transcriptional activator of antibiotic biosynthesis in Streptomyces.PLoS Biol. 2024 Mar 1;22(3):e3002528. doi: 10.1371/journal.pbio.3002528. eCollection 2024 Mar. PLoS Biol. 2024. PMID: 38427710 Free PMC article.
-
Recent advances in natural products exploitation in Streptomyces via synthetic biology.Eng Life Sci. 2019 Mar 12;19(6):452-462. doi: 10.1002/elsc.201800137. eCollection 2019 Jun. Eng Life Sci. 2019. PMID: 32625022 Free PMC article. Review.
-
The Transcriptional Regulator DhyR Positively Modulates Daptomycin Biosynthesis in Streptomyces roseosporus.Microb Biotechnol. 2025 Mar;18(3):e70110. doi: 10.1111/1751-7915.70110. Microb Biotechnol. 2025. PMID: 40025688 Free PMC article.
-
Coordinated regulation for nature products discovery and overproduction in Streptomyces.Synth Syst Biotechnol. 2020 Apr 20;5(2):49-58. doi: 10.1016/j.synbio.2020.04.002. eCollection 2020 Jun. Synth Syst Biotechnol. 2020. PMID: 32346621 Free PMC article.
-
Biosynthesis of Polyketides in Streptomyces.Microorganisms. 2019 May 6;7(5):124. doi: 10.3390/microorganisms7050124. Microorganisms. 2019. PMID: 31064143 Free PMC article. Review.
References
-
- Menéndez N, Nur-e-Alam M, Brana AF, Rohr J, Salas JA, Méndez C. Biosynthesis of the antitumor chromomycin A3 in Streptomyces griseus. analysis of the gene cluster and rational design of novel chromomycin analogs. Chem Biol. 2004;11(1):21–32. - PubMed
-
- Skarbek JD, Speedie MK. Antitumor antibiotics of the aureolic acid group: chromomycin A3, mithramycin A, and olivomycin A. In Antitumor Compounds of Natural Origin. (A. Aszalos, ed.). Chem Biochemistry. Boca-Raton: CRC Press; 1981;I:191–235.
-
- Bianchi N, Osti F, Rutigliano C, Corradini FG, Borsetti E, Tomassetti M, Mischiati C, Feriotto G, Gambari R. The DNA-binding drugs mithramycin and chromomycin are powerful inducers of erythroid differentiation of human K562 cells. Br J Haematol. 1999;104(2):258–265. doi: 10.1046/j.1365-2141.1999.01173.x. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

